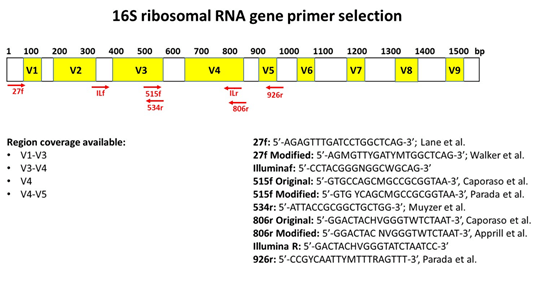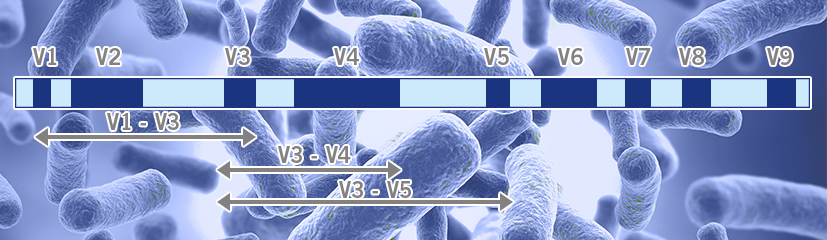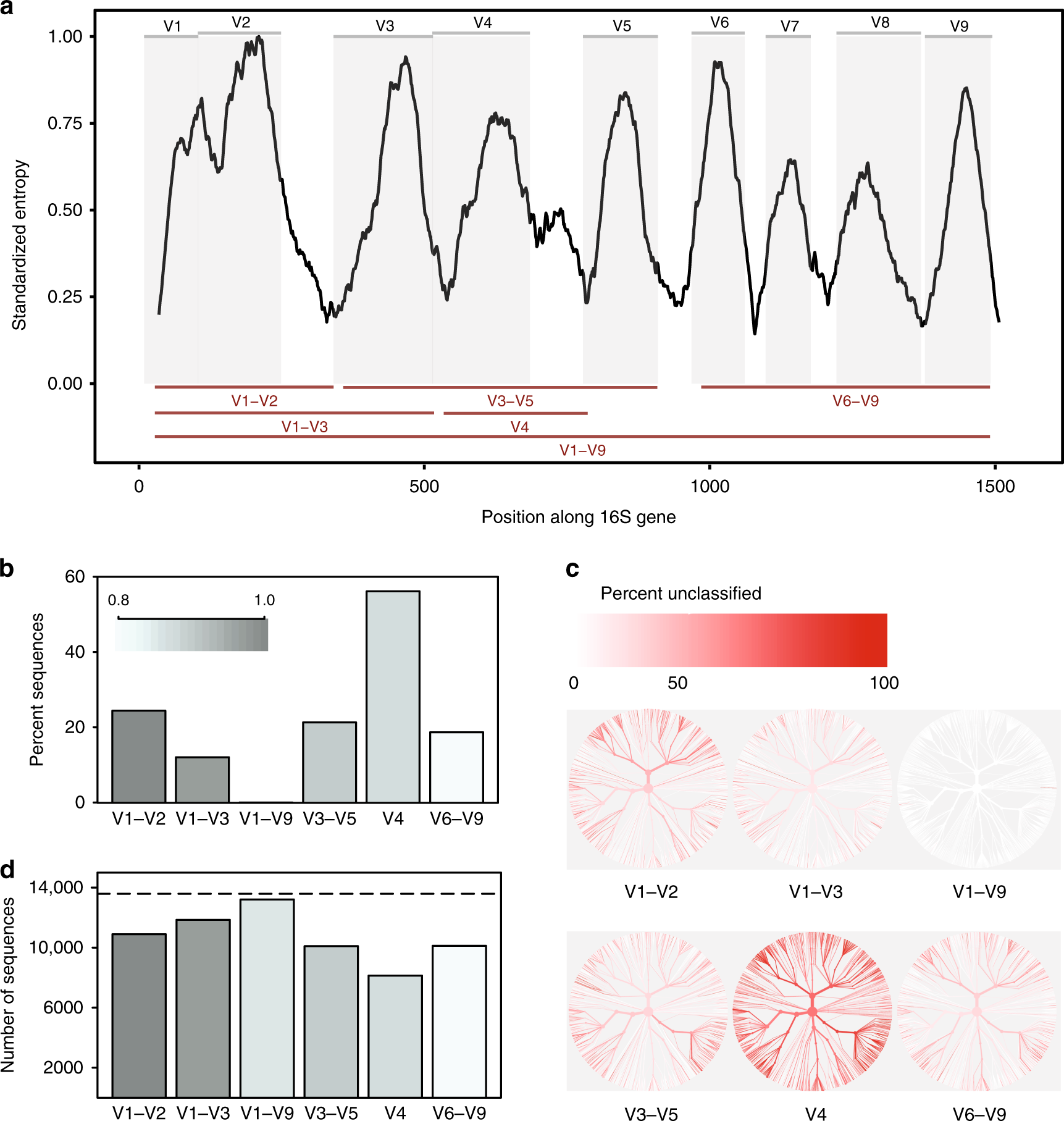
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications

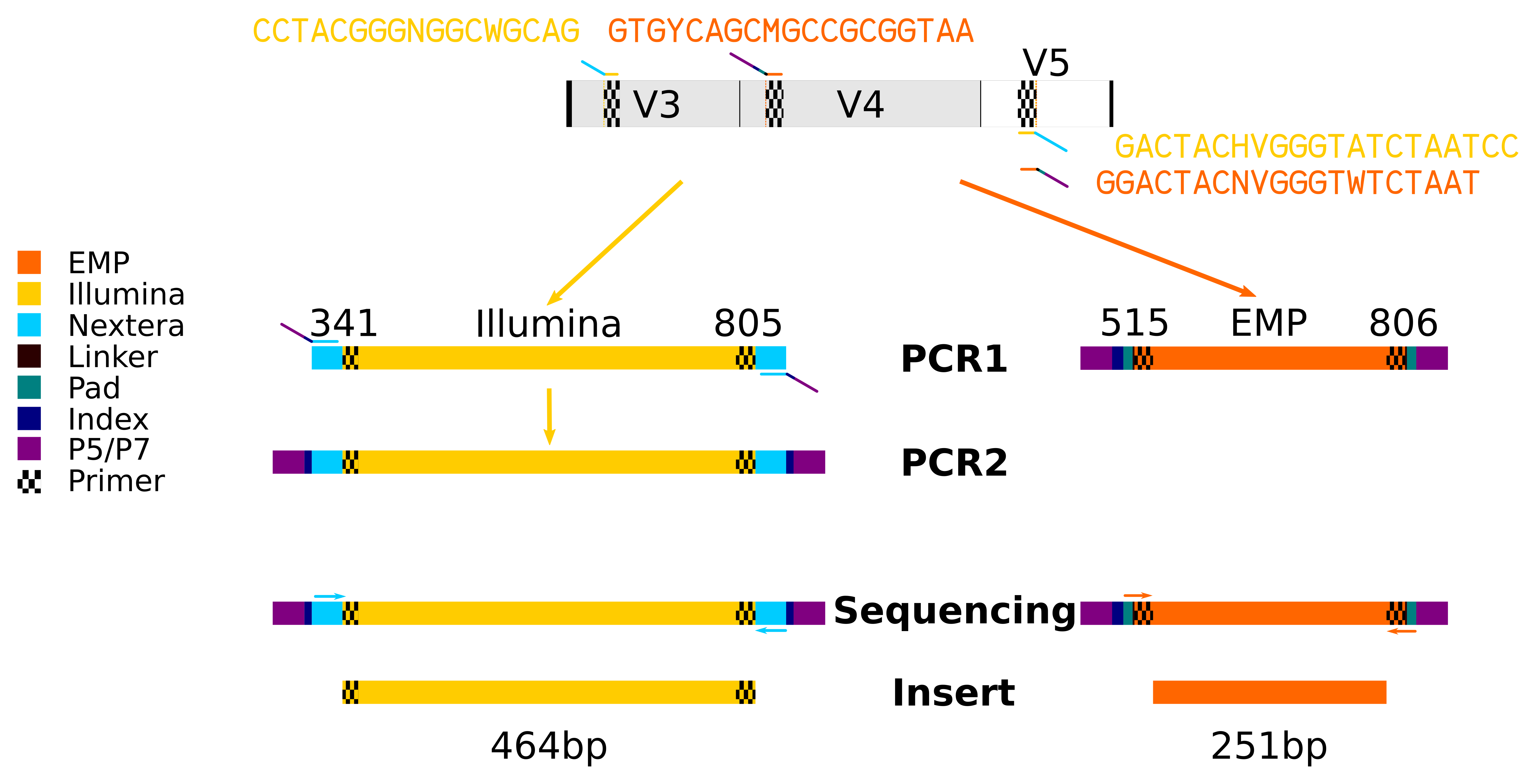
Evaluation of Compatibility of 16S rRNA V3V4 and V4 Amplicon Libraries for Clinical Microbiome Profiling | bioRxiv

Reproducibility and repeatability of six high-throughput 16S rDNA sequencing protocols for microbiota profiling - ScienceDirect

Evaluation of Compatibility of 16S rRNA V3V4 and V4 Amplicon Libraries for Clinical Microbiome Profiling | bioRxiv
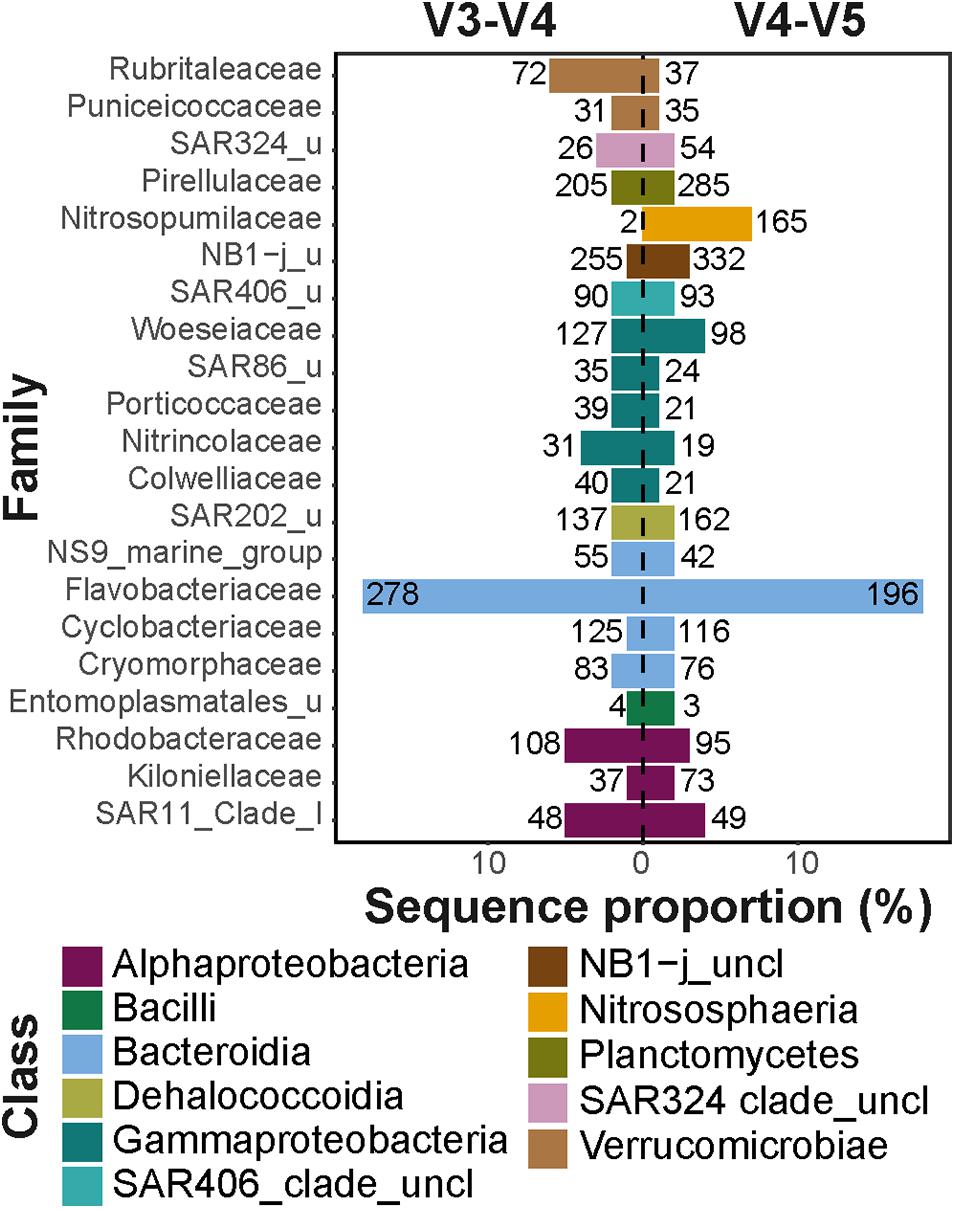
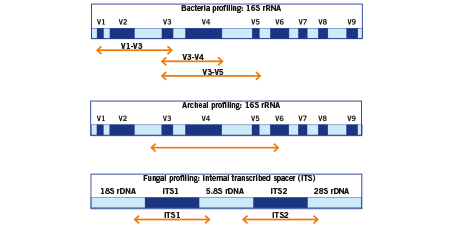
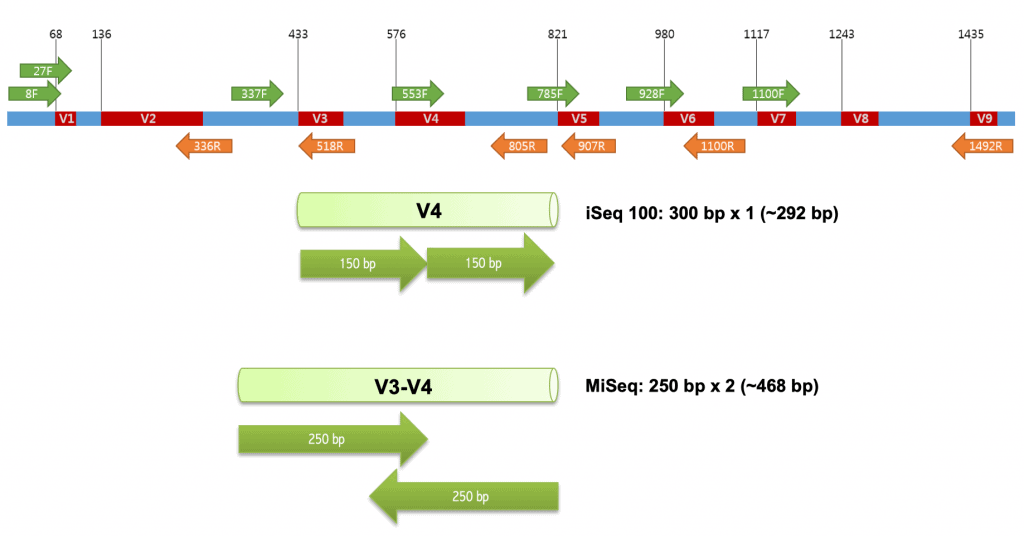
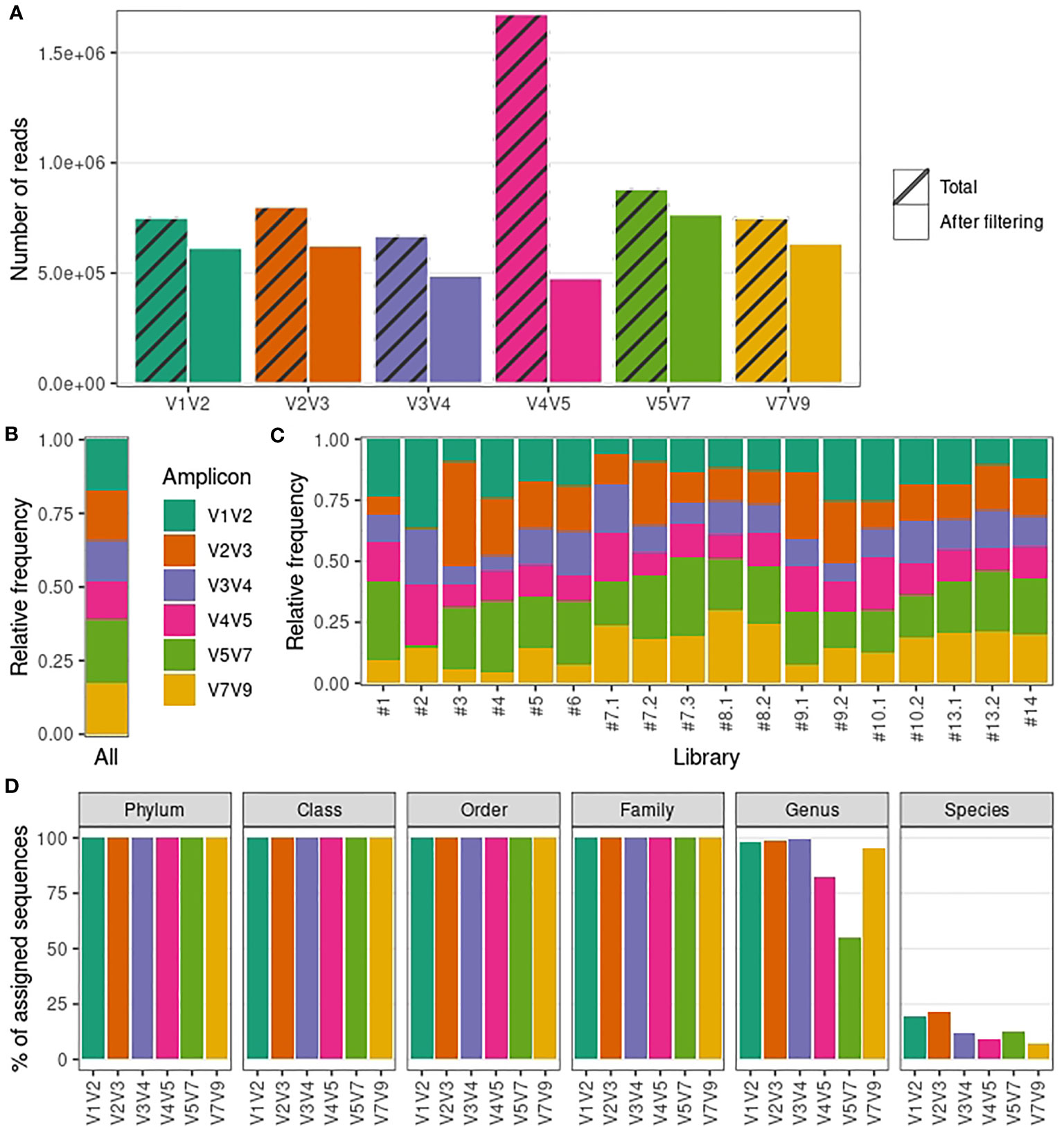
Frontiers | Comparison of Two 16S rRNA Primers (V3–V4 and V4–V5) for Studies of Arctic Microbial Communities

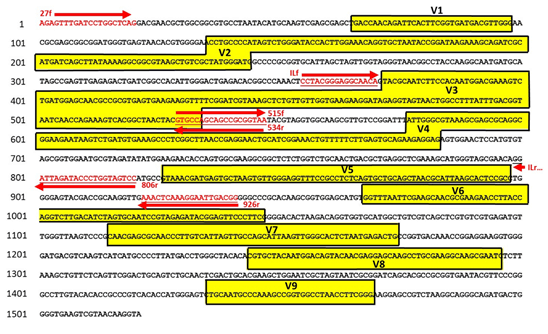
Modification of the 16S V3-V4 reverse primer reduced co-amplification... | Download Scientific Diagram
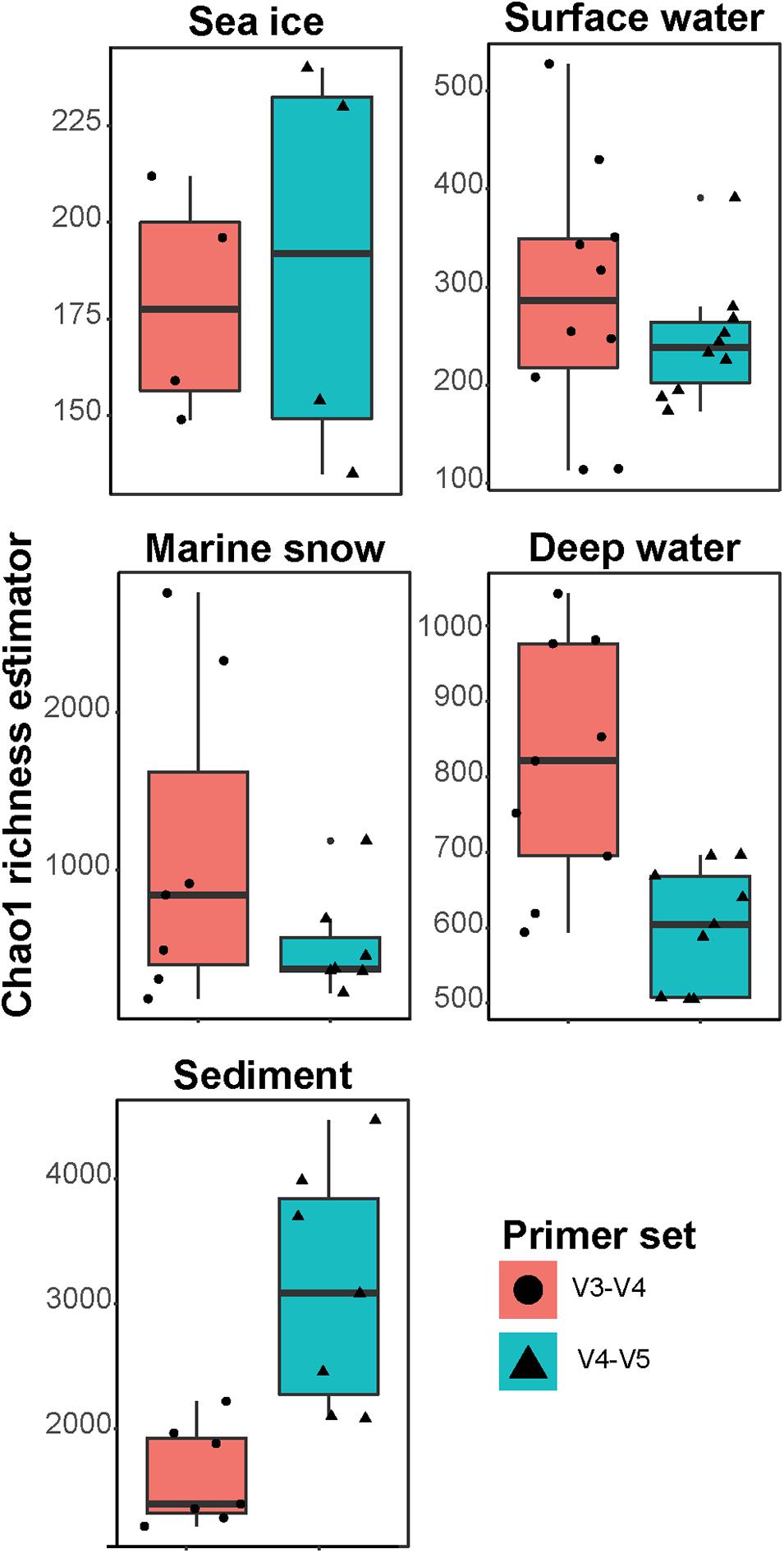
Frontiers | Comparison of Two 16S rRNA Primers (V3–V4 and V4–V5) for Studies of Arctic Microbial Communities

An optimized 16S rRNA sequencing protocol for vaginal microbiome to avoid biased abundance estimation | bioRxiv

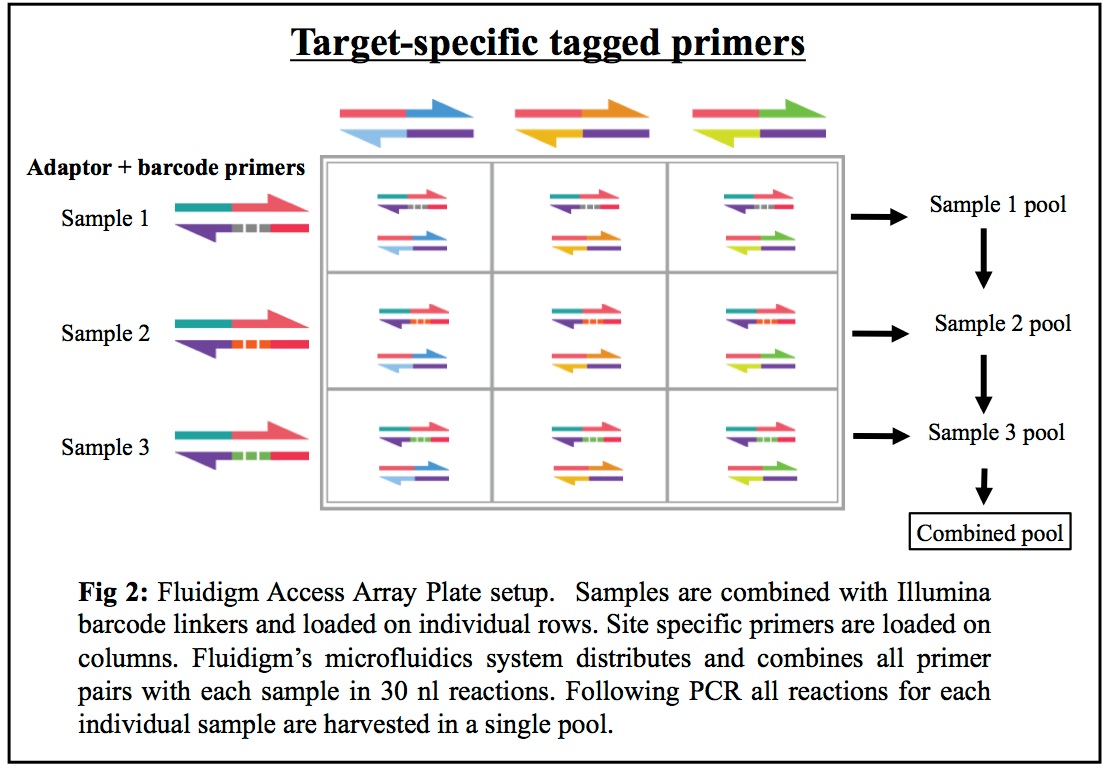
Schematic diagram of 16S rDNA sequencing workflow for both Run 1 and... | Download Scientific Diagram

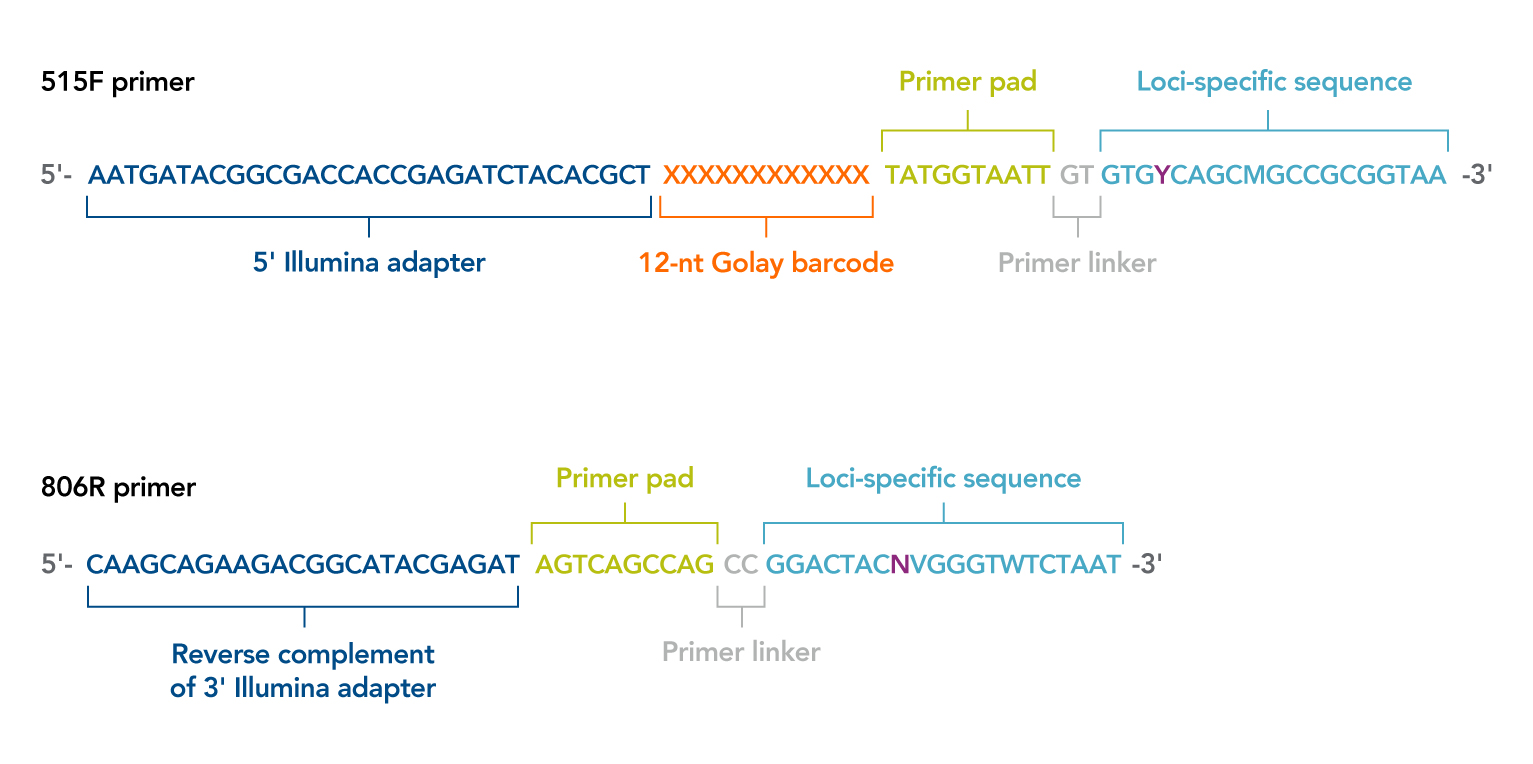
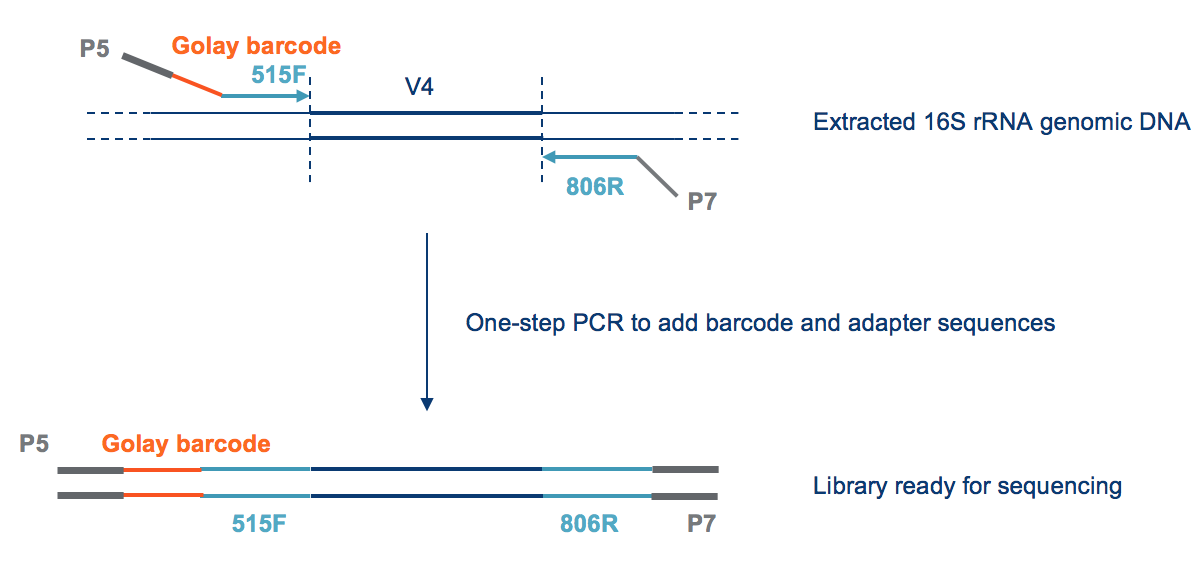
Better skin microbiome analyses using new 16S V4 region primers developed by Microbiome Insights' scientific team — Microbiome Insights