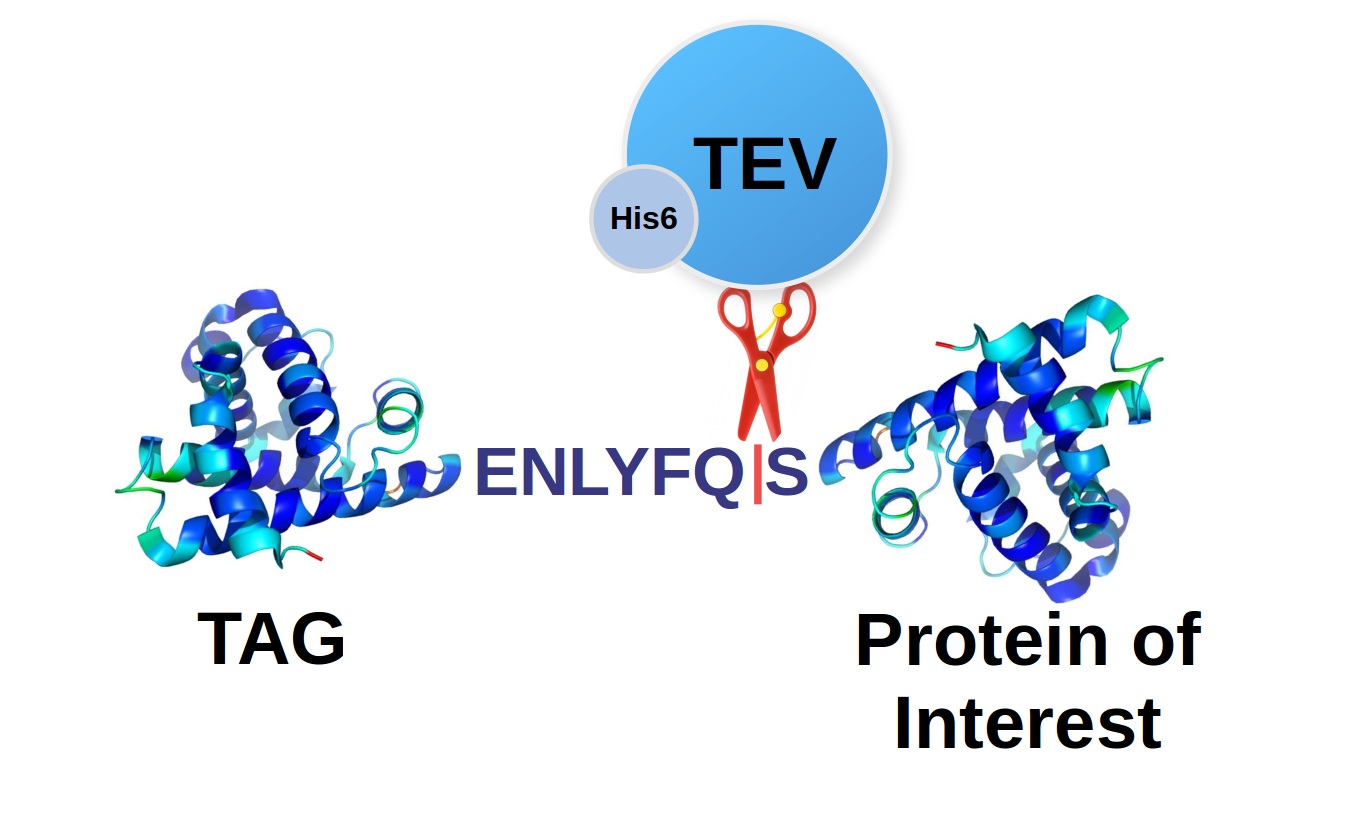
Improved yield, stability, and cleavage reaction of a novel tobacco etch virus protease mutant | SpringerLink
A novel family of expression vectors with multiple affinity tags for wheat germ cell-free protein expression | BMC Biotechnology | Full Text
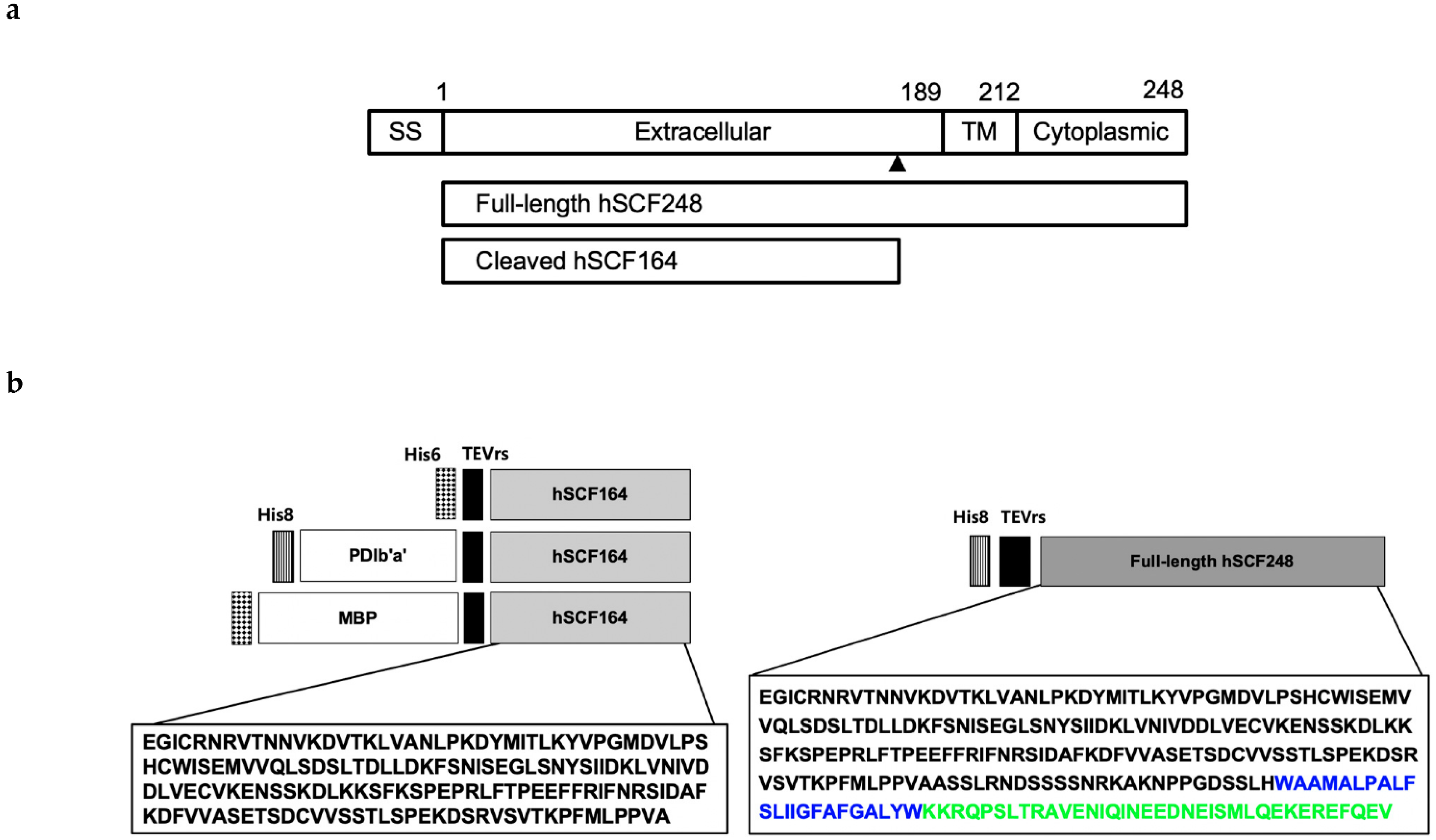
IJMS | Free Full-Text | Novel Bacterial Production of Two Different Bioactive Forms of Human Stem-Cell Factor

Deep profiling of protease substrate specificity enabled by dual random and scanned human proteome substrate phage libraries | PNAS

Three-Amino Acid Spacing of Presenilin Endoproteolysis Suggests a General Stepwise Cleavage of γ-Secretase-Mediated Intramembrane Proteolysis | Journal of Neuroscience

YESS 2.0, a Tunable Platform for Enzyme Evolution, Yields Highly Active TEV Protease Variants | ACS Synthetic Biology

Three-Amino Acid Spacing of Presenilin Endoproteolysis Suggests a General Stepwise Cleavage of γ-Secretase-Mediated Intramembrane Proteolysis | Journal of Neuroscience
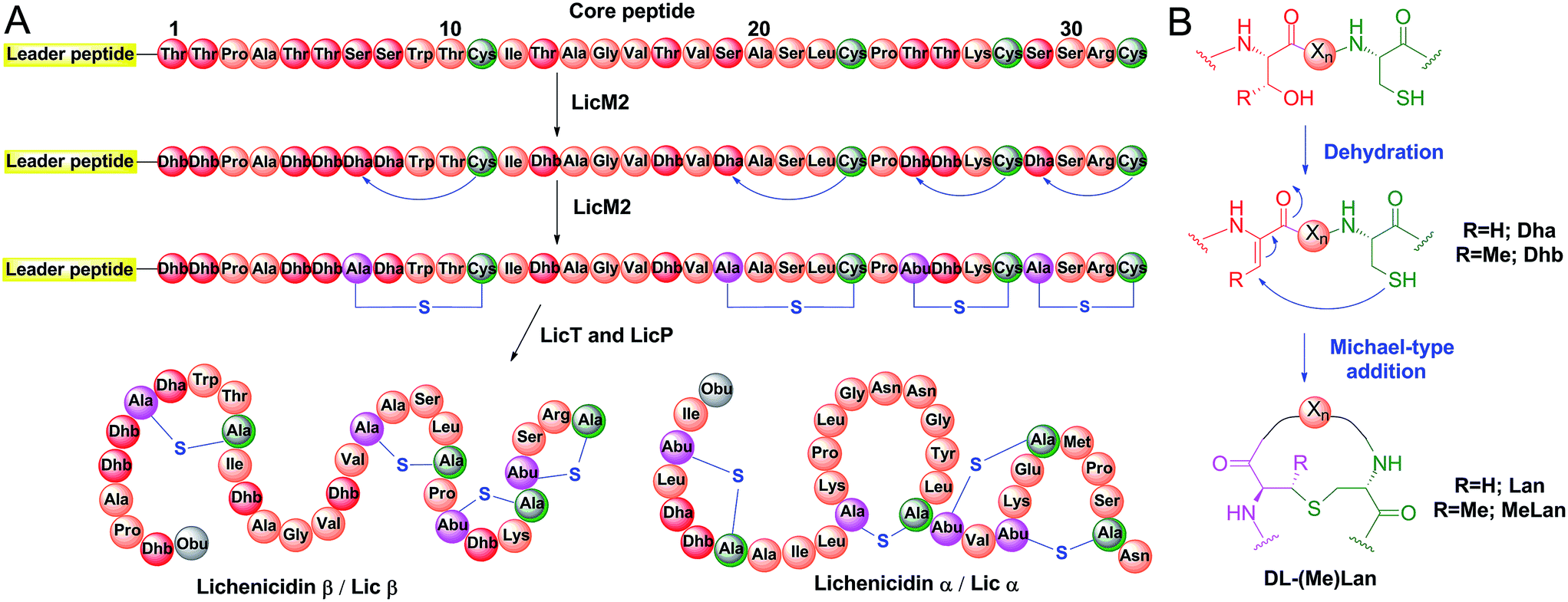
Applications of the class II lanthipeptide protease LicP for sequence-specific, traceless peptide bond cleavage - Chemical Science (RSC Publishing) DOI:10.1039/C5SC02329G

SDS-PAGE analysis of TEV protease cleavage efficiency. (A) 0.94 mg and... | Download Scientific Diagram
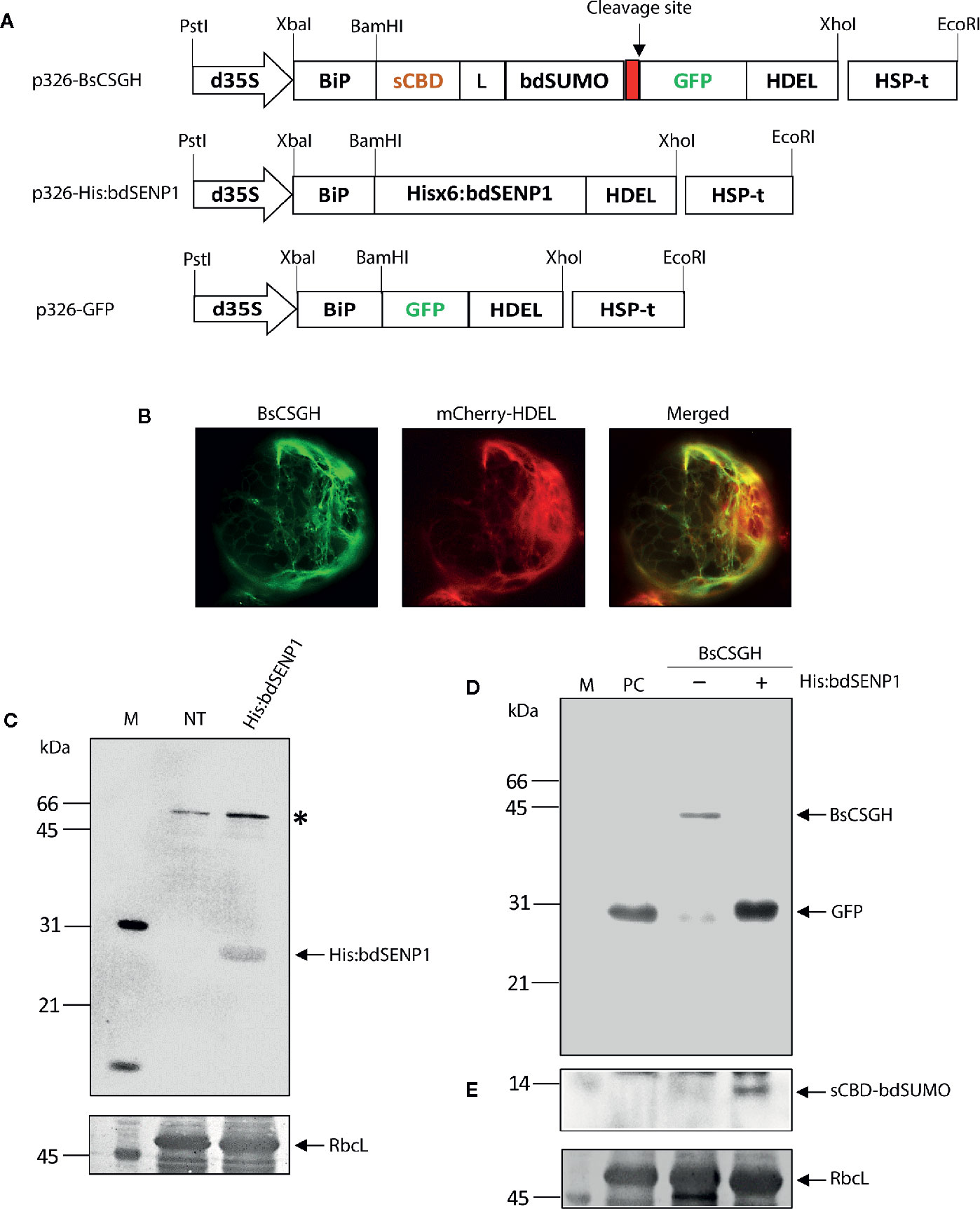
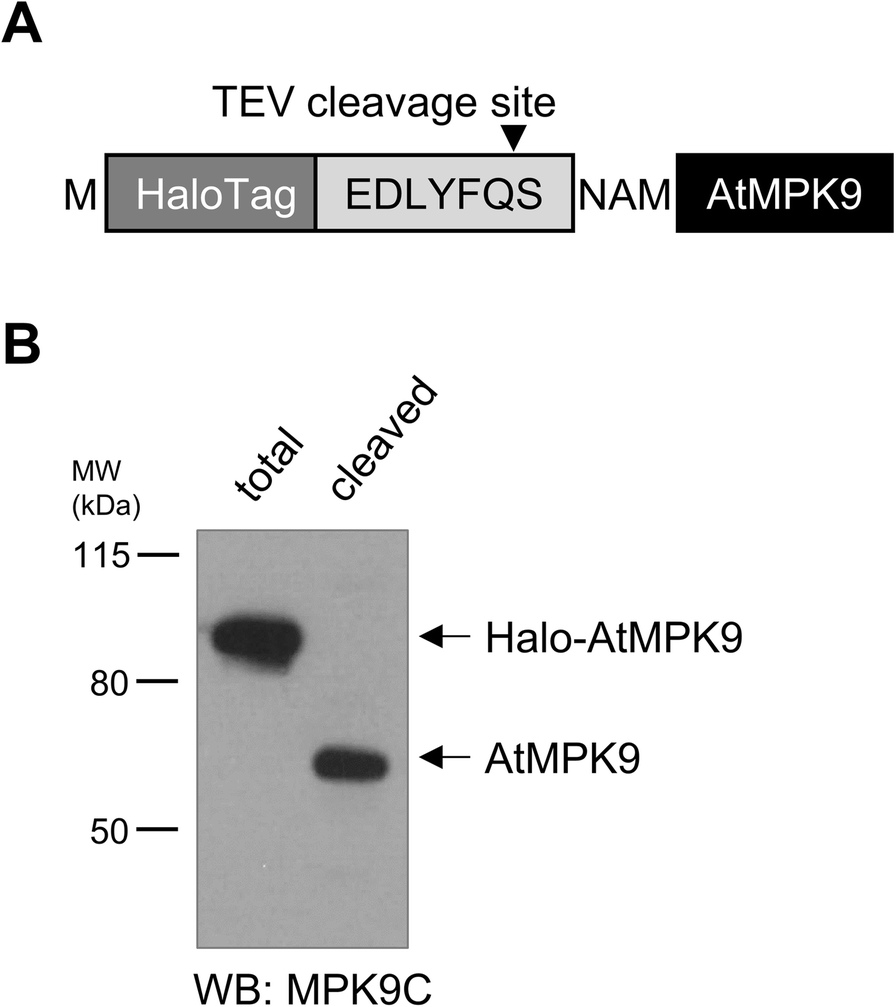
Frontiers | In Vivo Removal of N-Terminal Fusion Domains From Recombinant Target Proteins Produced in Nicotiana benthamiana

Schematic presentation of protease TEV cleavage site, V8 cleavage site... | Download Scientific Diagram
Highly efficient soluble expression, purification and characterization of recombinant Aβ42 from Escherichia coli

Going native: Complete removal of protein purification affinity tags by simple modification of existing tags and proteases - ScienceDirect

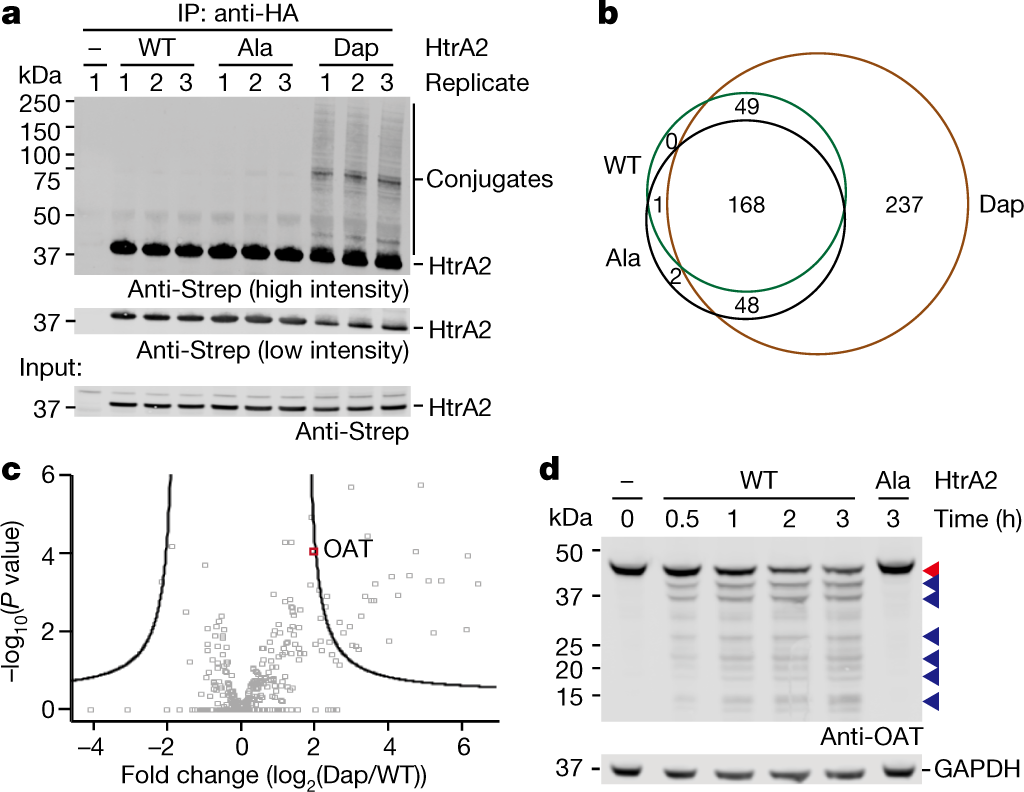
Global Sequencing of Proteolytic Cleavage Sites in Apoptosis by Specific Labeling of Protein N Termini: Cell

Expression strategy of the CBD-lunasin fusion protein and subsequent... | Download Scientific Diagram