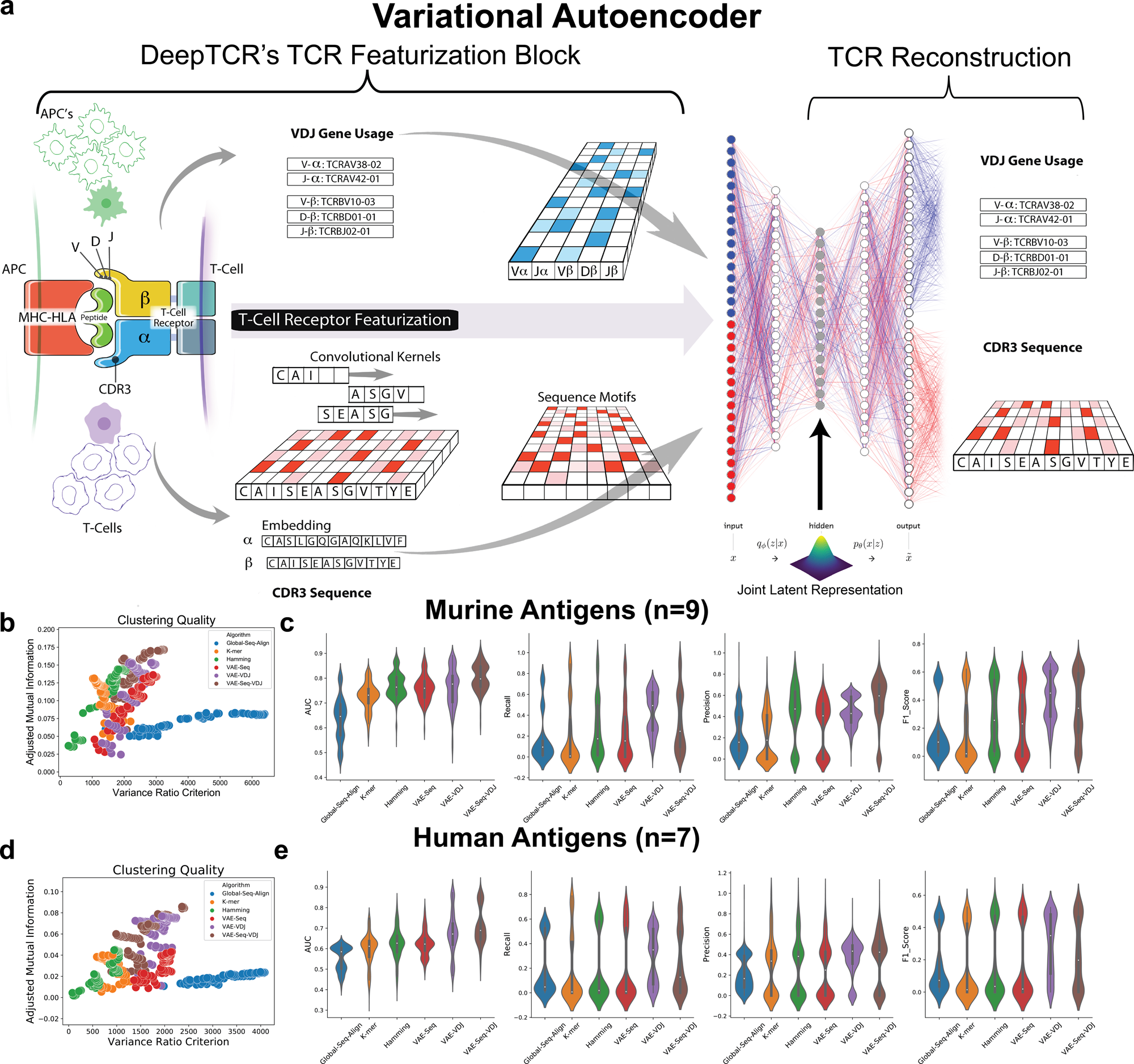
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications

Recent advances in T-cell receptor repertoire analysis: Bridging the gap with multimodal single-cell RNA sequencing - ScienceDirect

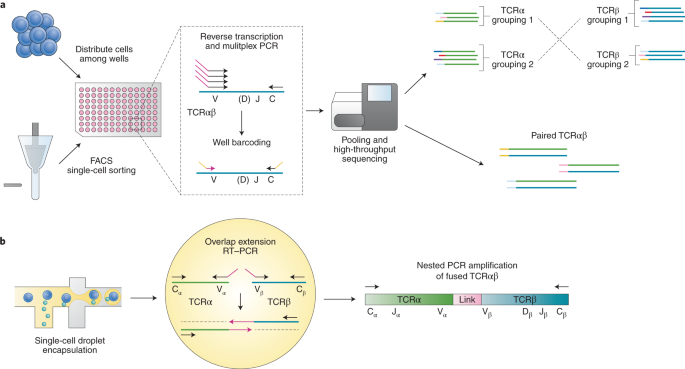
Recent advances in T-cell receptor repertoire analysis: Bridging the gap with multimodal single-cell RNA sequencing - ScienceDirect
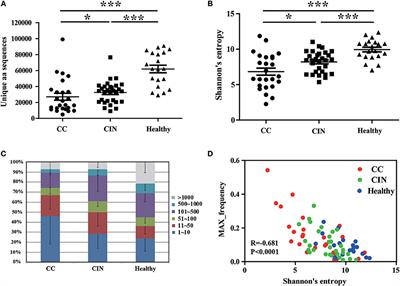
Frontiers | TCR Repertoire as a Novel Indicator for Immune Monitoring and Prognosis Assessment of Patients With Cervical Cancer

pyTCR: a comprehensive and scalable platform for TCR-Seq data analysis to facilitate reproducibility and rigor of immunogenomics research | medRxiv
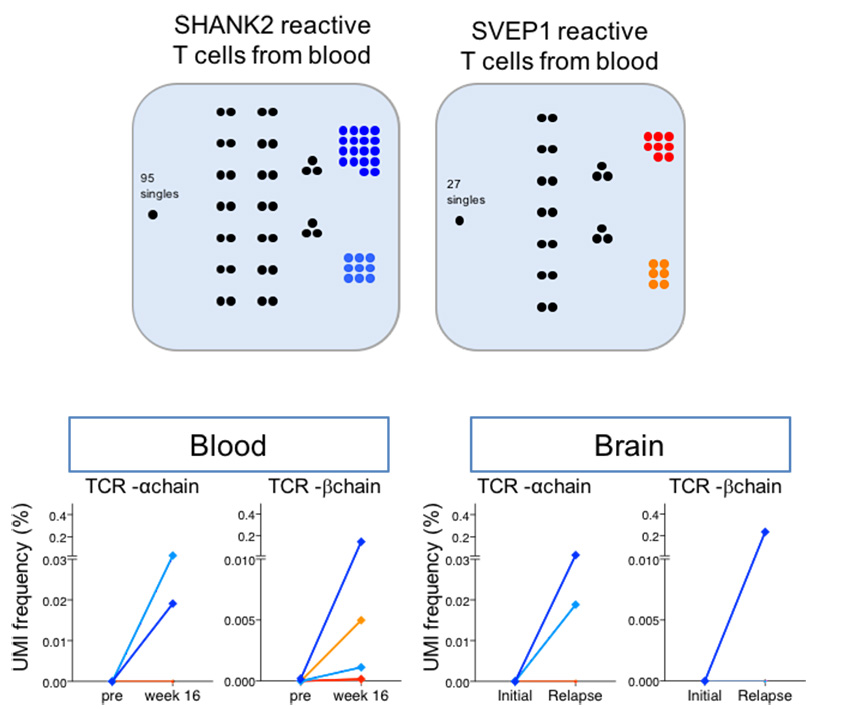
Detecting T cell receptors involved in immune responses from single repertoire snapshots | PLOS Biology

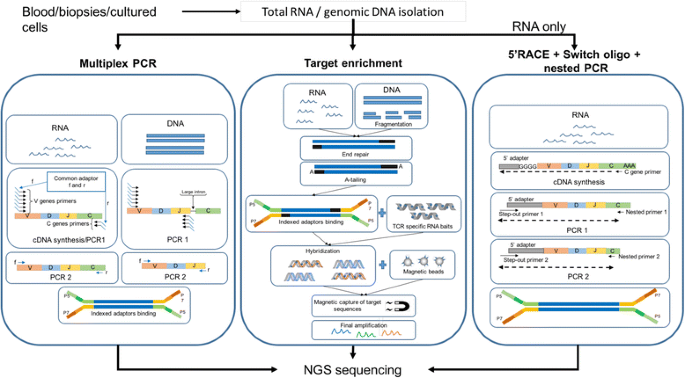
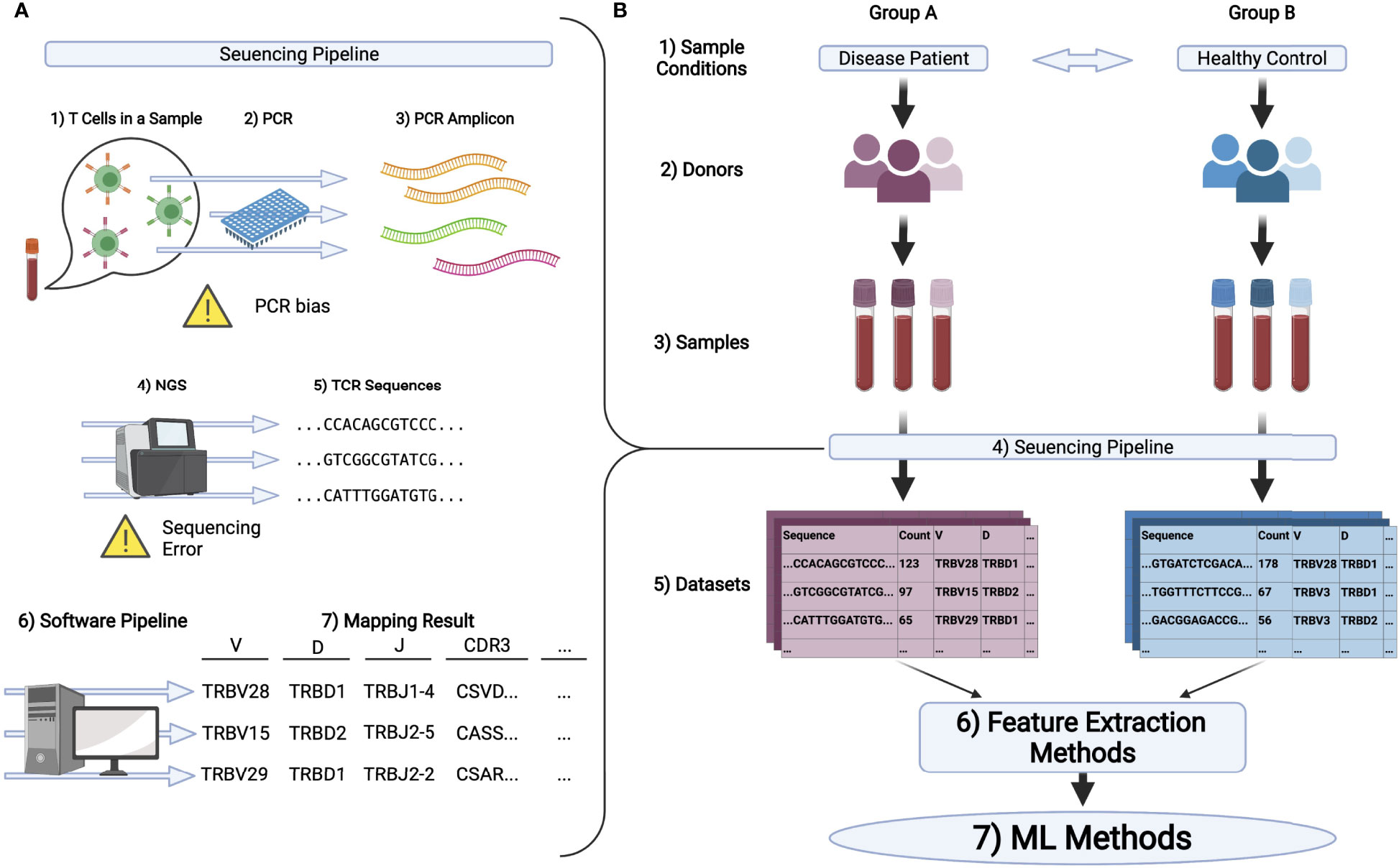
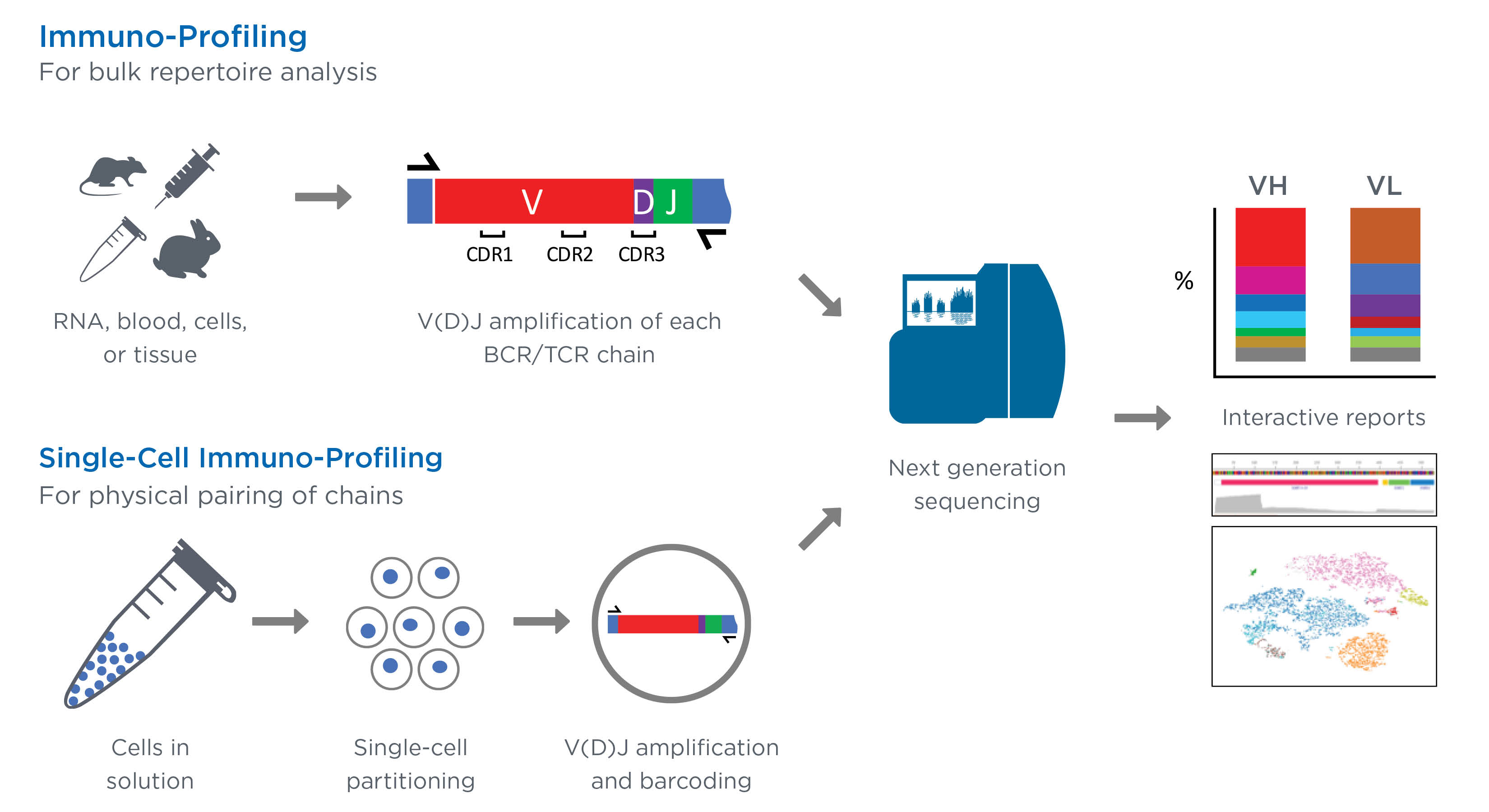
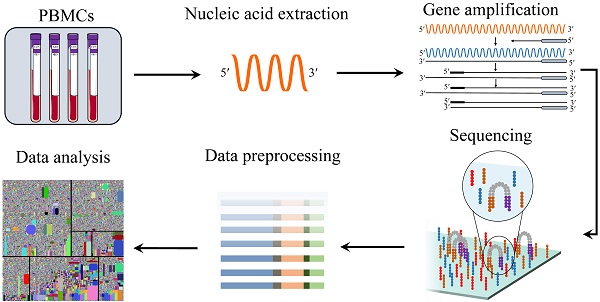
General workflow for TCR repertoire sequencing and analysis. From bulk... | Download Scientific Diagram

A framework for highly multiplexed dextramer mapping and prediction of T cell receptor sequences to antigen specificity | Science Advances
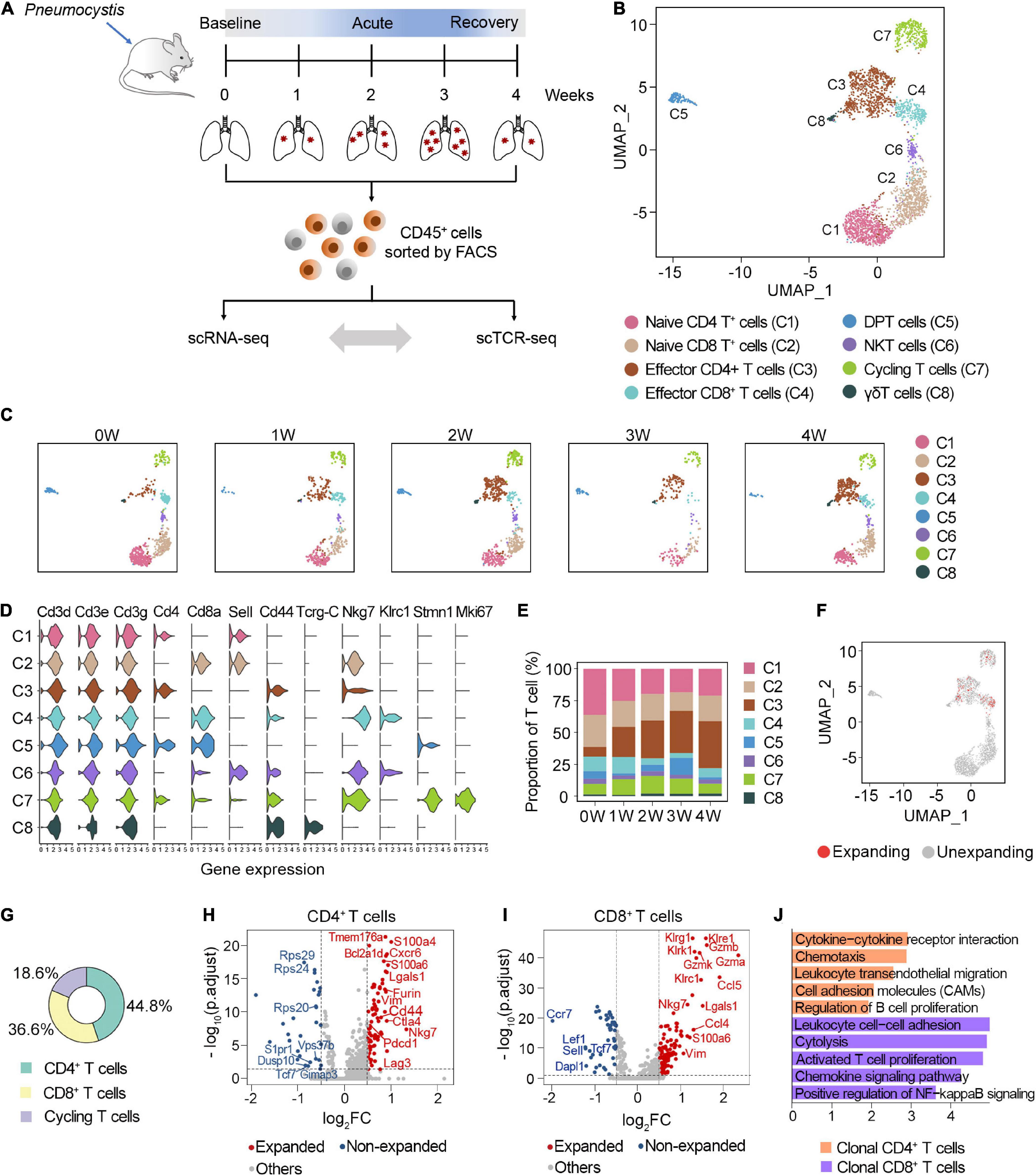
Frontiers | Single-Cell TCR Sequencing Reveals the Dynamics of T Cell Repertoire Profiling During Pneumocystis Infection

DeepTCR: a deep learning framework for understanding T-cell receptor sequence signatures within complex T-cell repertoires | bioRxiv
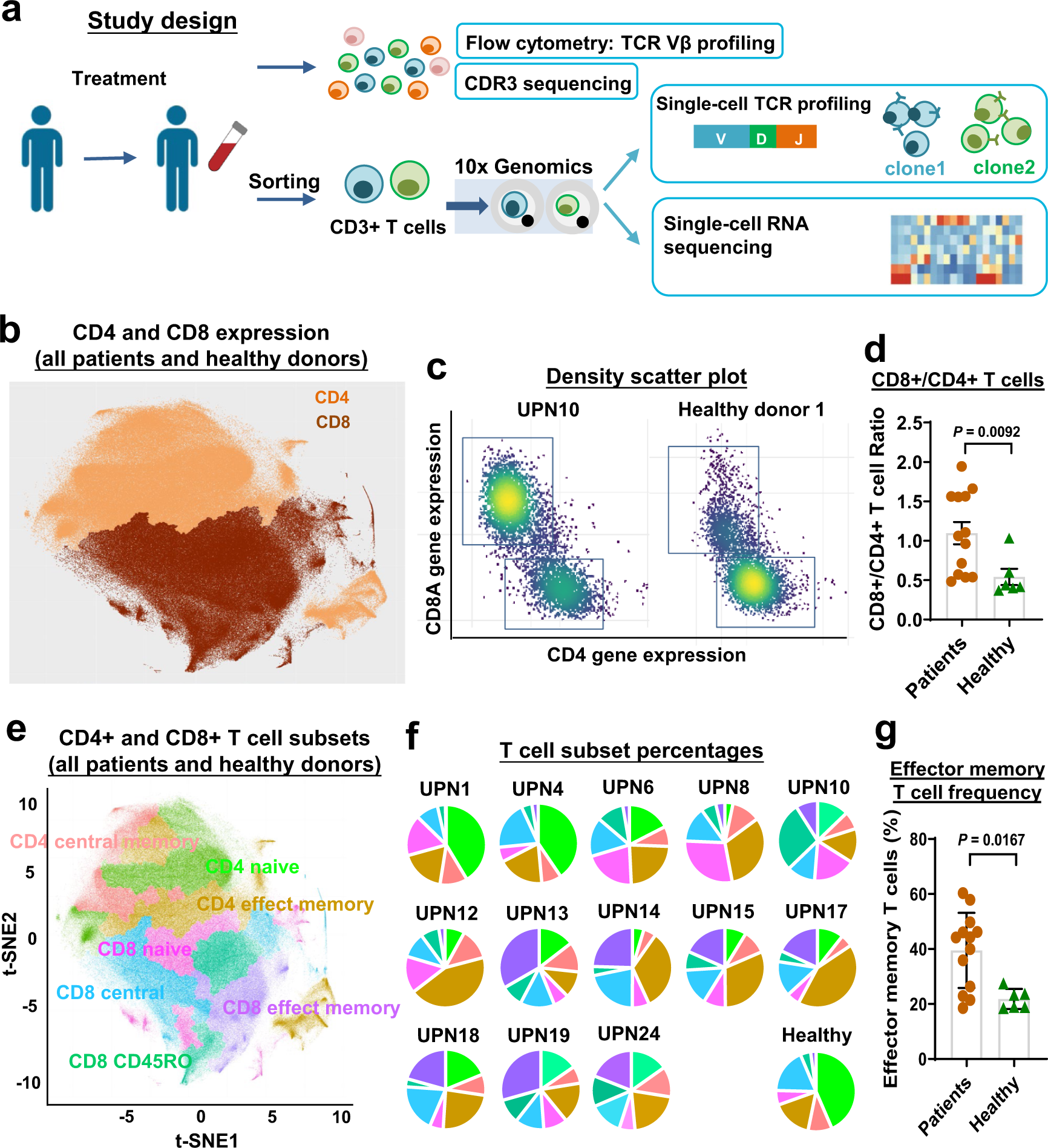
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells | Nature Communications