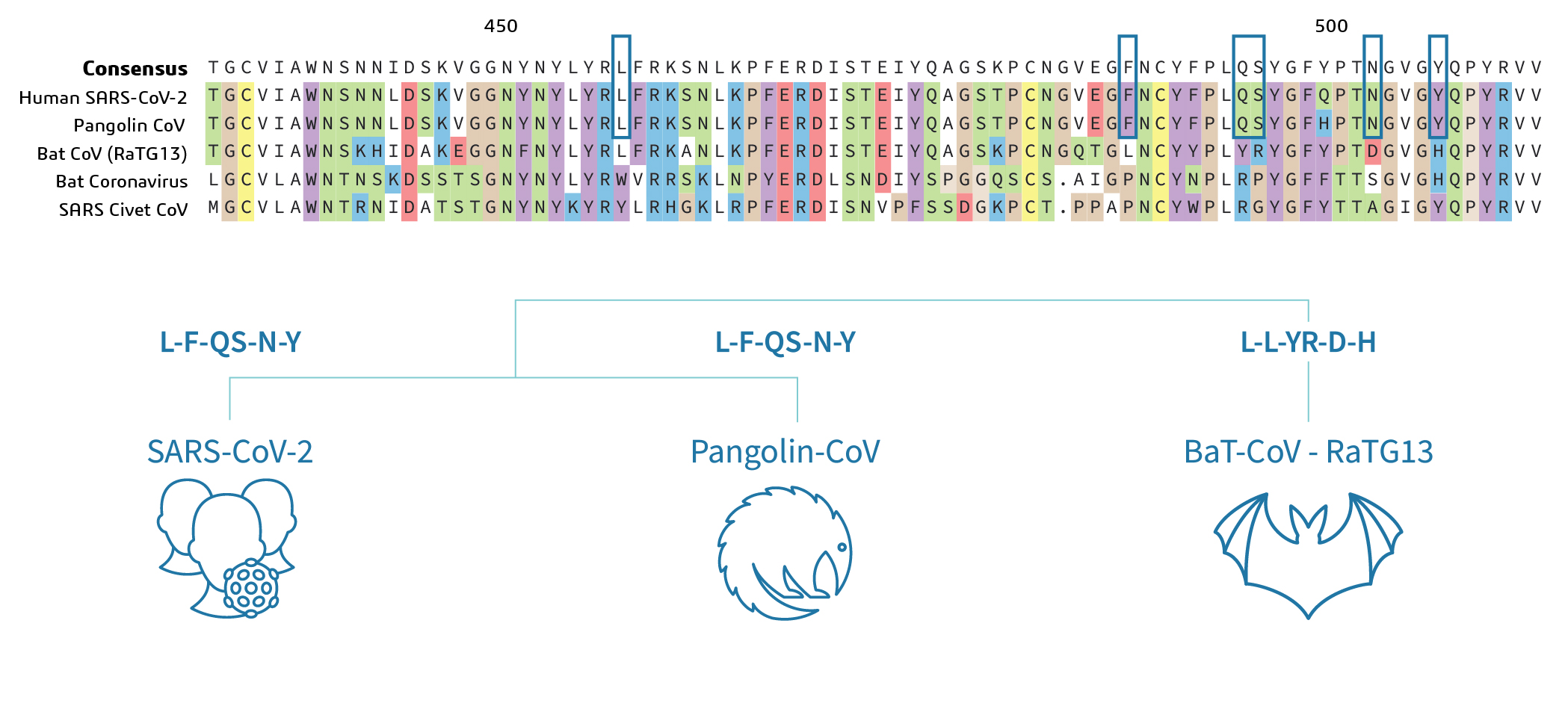
Tracking the amino acid changes of spike proteins across diverse host species of severe acute respiratory syndrome coronavirus 2 - ScienceDirect
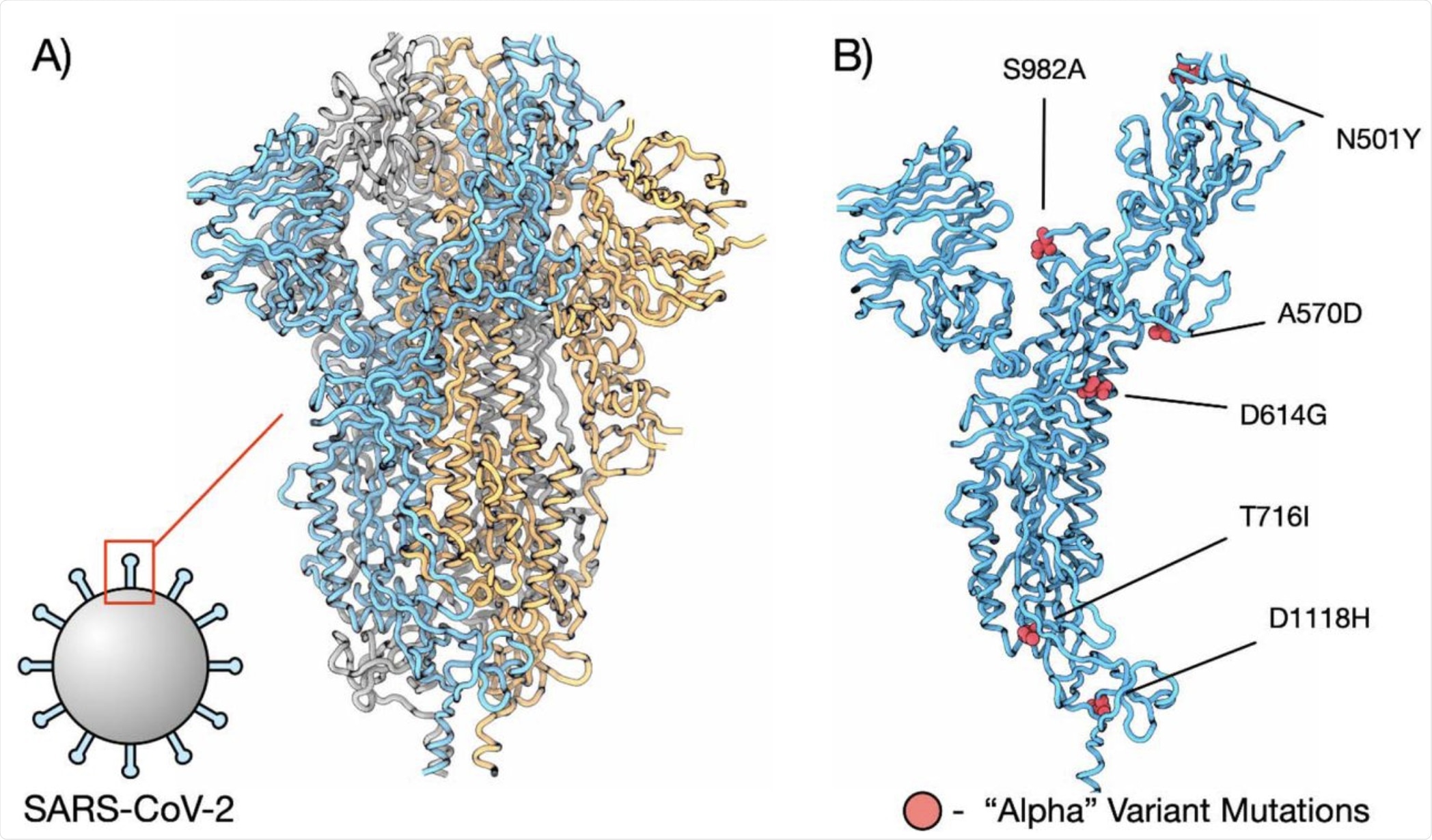
Mutations in the spike proteins of SARS-CoV-2 select for amino acid changes, increasing protein stability

Wide variabilities identified among spike proteins of SARS Cov2 globally-dominant variant identified | bioRxiv

Identification of Two Critical Amino Acid Residues of the Severe Acute Respiratory Syndrome Coronavirus Spike Protein for Its Variation in Zoonotic Tropism Transition via a Double Substitution Strategy - ScienceDirect
Implications of the Mutations in the Spike Protein of the Omicron Variant of Concern (VoC) of SARS-CoV-2 – Signature Science

The “LLQY” Motif on SARS-CoV-2 Spike Protein Affects S Incorporation into Virus Particles | Journal of Virology
nCoV-2019 Spike Protein Receptor Binding Domain Shares High Amino Acid Identity With a Coronavirus Recovered from a Pangolin Viral Metagenomic Dataset - nCoV-2019 Evolutionary History - Virological
Molecular epidemiology of SARS-CoV-2 clusters caused by asymptomatic cases in Anhui Province, China | BMC Infectious Diseases | Full Text

SARS-CoV-2 Omicron variant: Antibody evasion and cryo-EM structure of spike protein–ACE2 complex | Science

Linear B-cell epitopes in the spike and nucleocapsid proteins as markers of SARS-CoV-2 exposure and disease severity - eBioMedicine

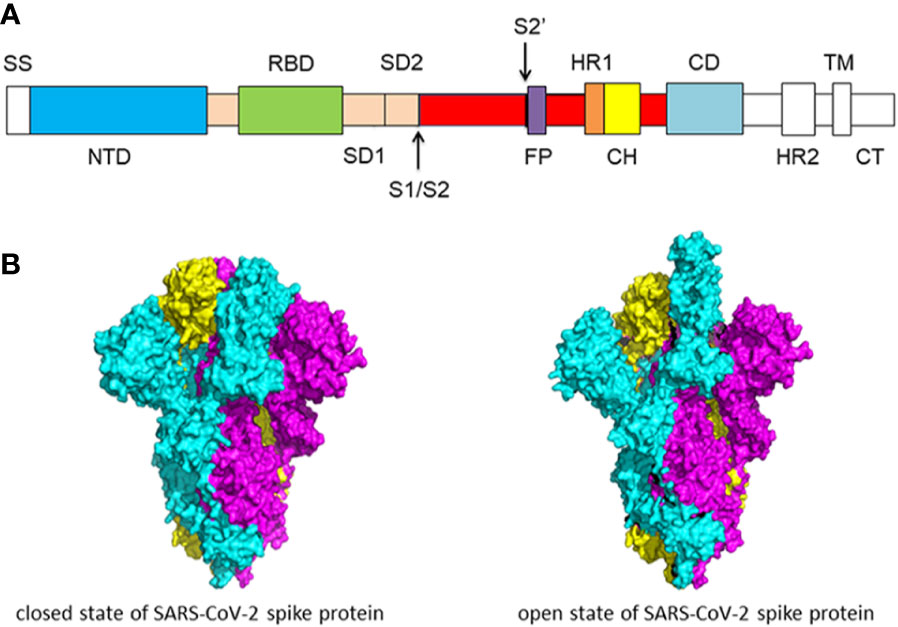
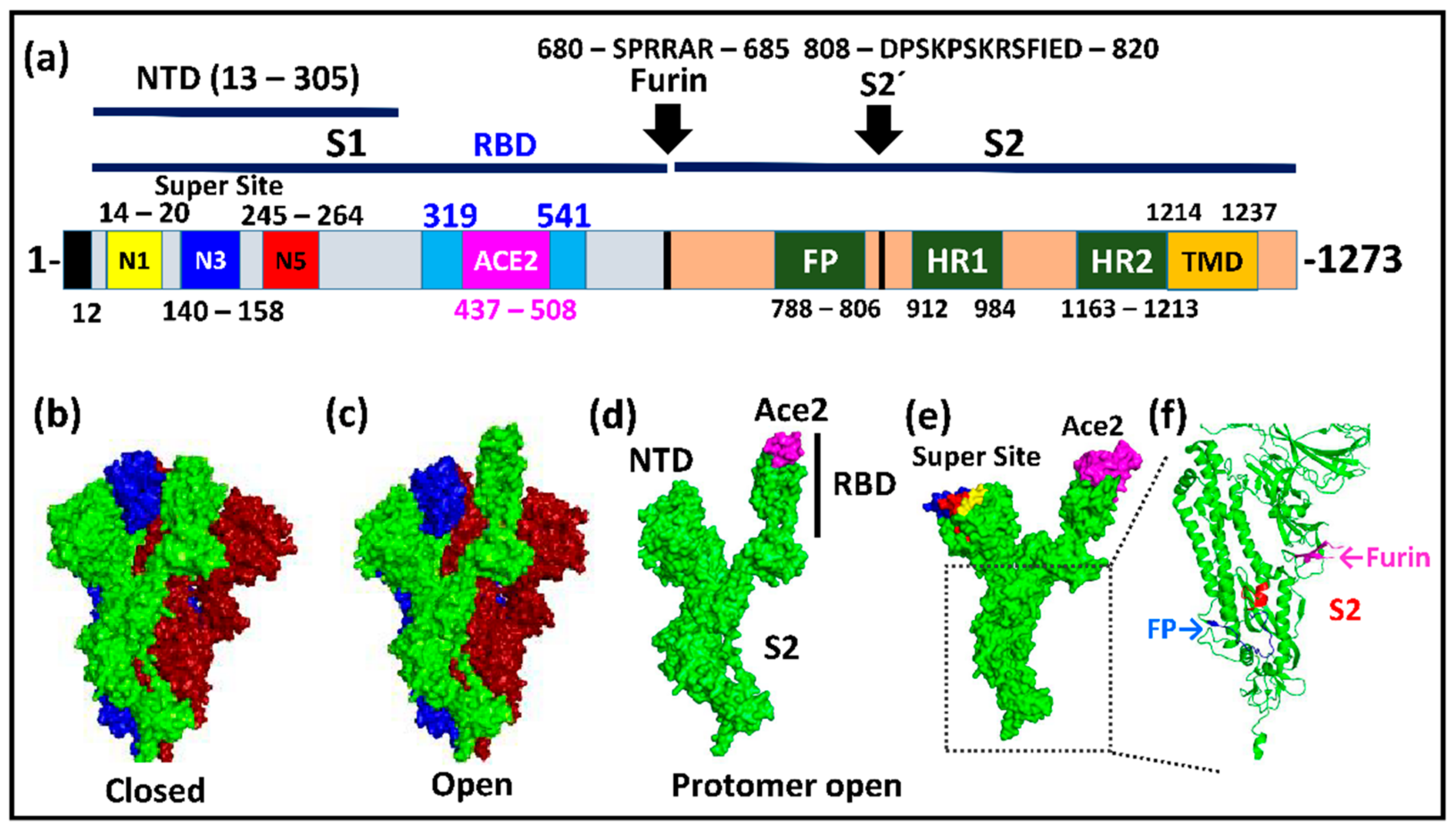
Molecular biology of the SARs‐CoV‐2 spike protein: A review of current knowledge - Zhu - 2021 - Journal of Medical Virology - Wiley Online Library
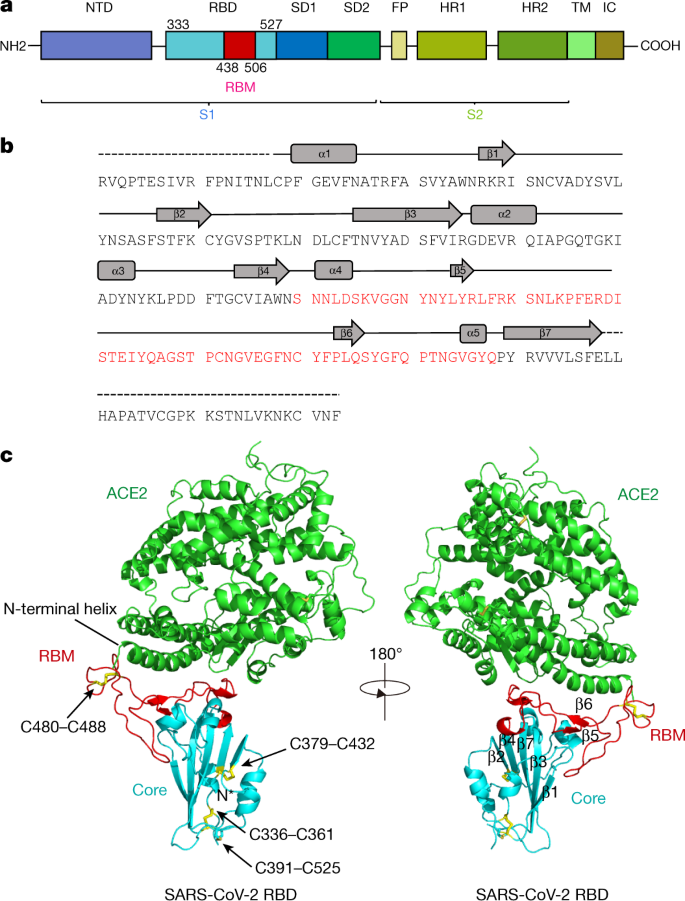
Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine | Cellular & Molecular Immunology
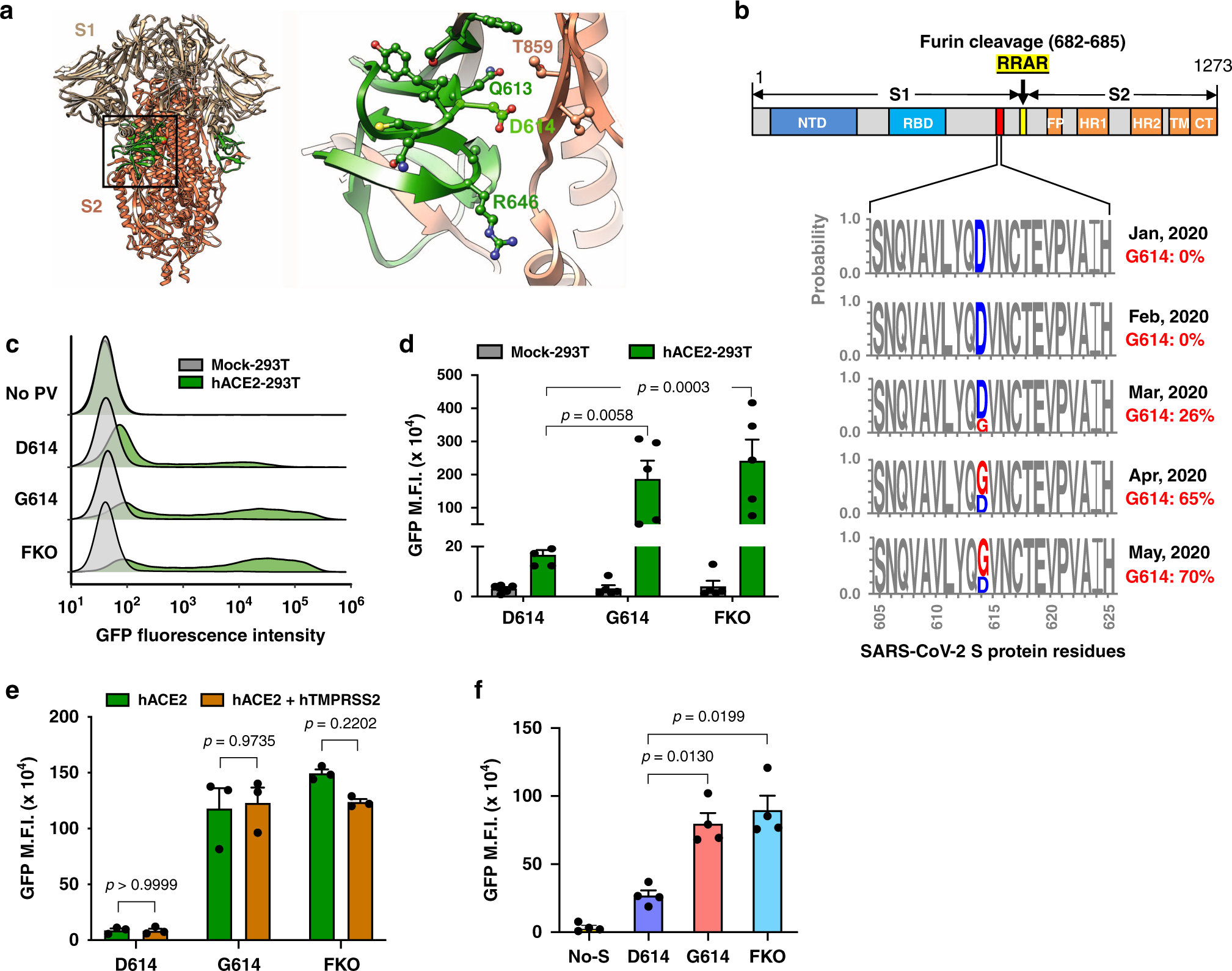
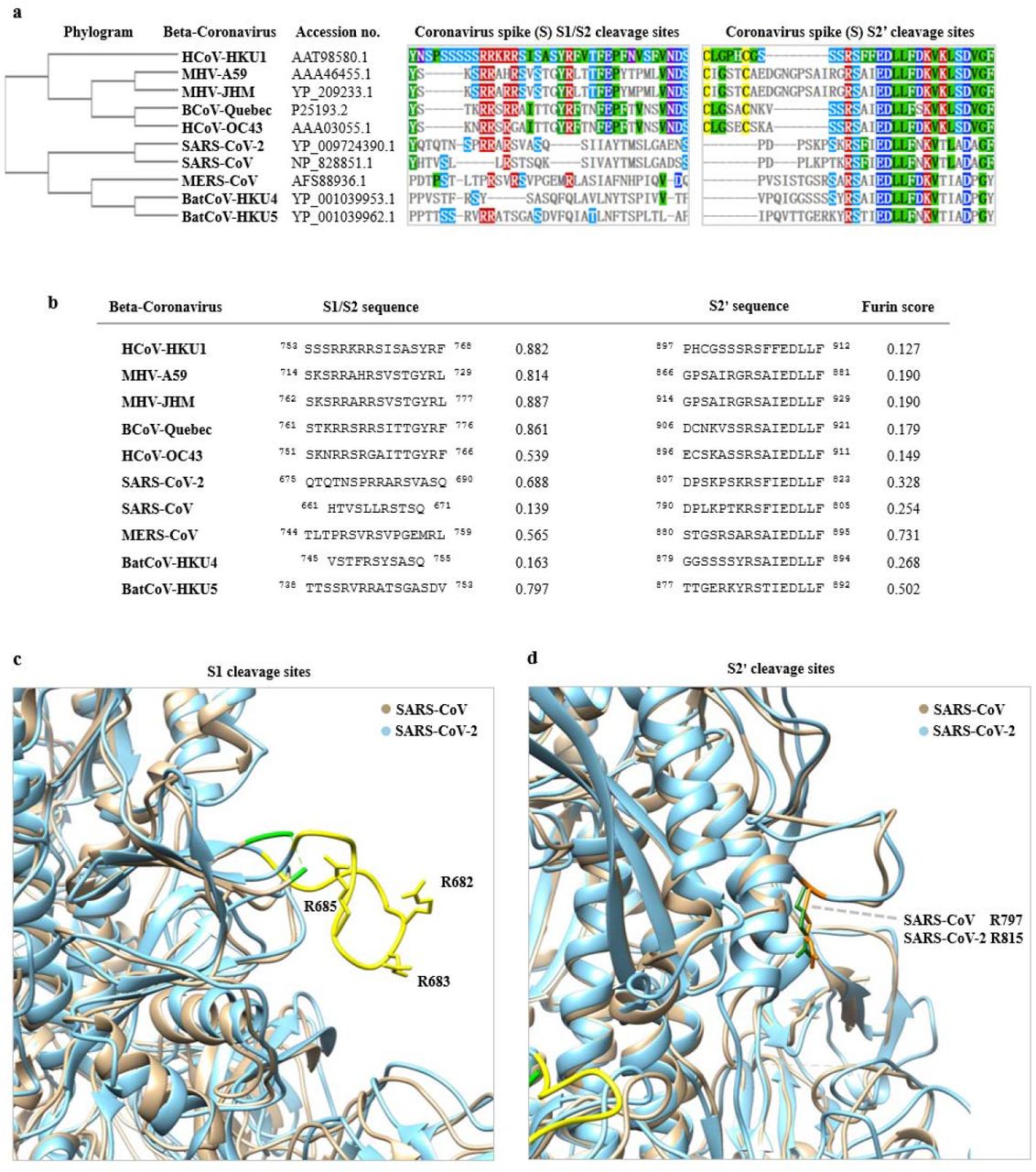
The role of furin cleavage site in SARS-CoV-2 spike protein-mediated membrane fusion in the presence or absence of trypsin | Signal Transduction and Targeted Therapy
Spike protein mutations in novel SARS-CoV-2 'variants of concern' commonly occur in or near indels - Virological