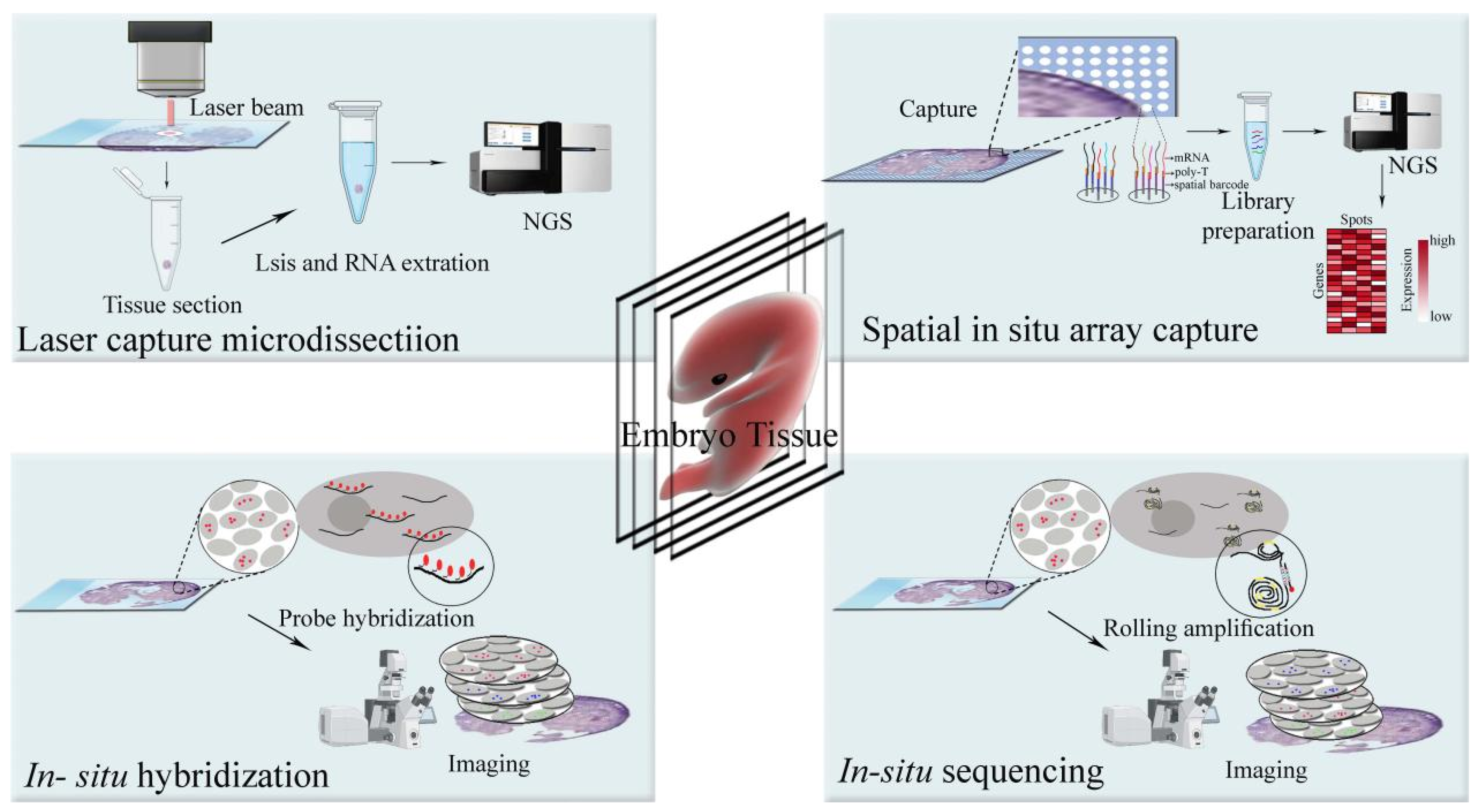
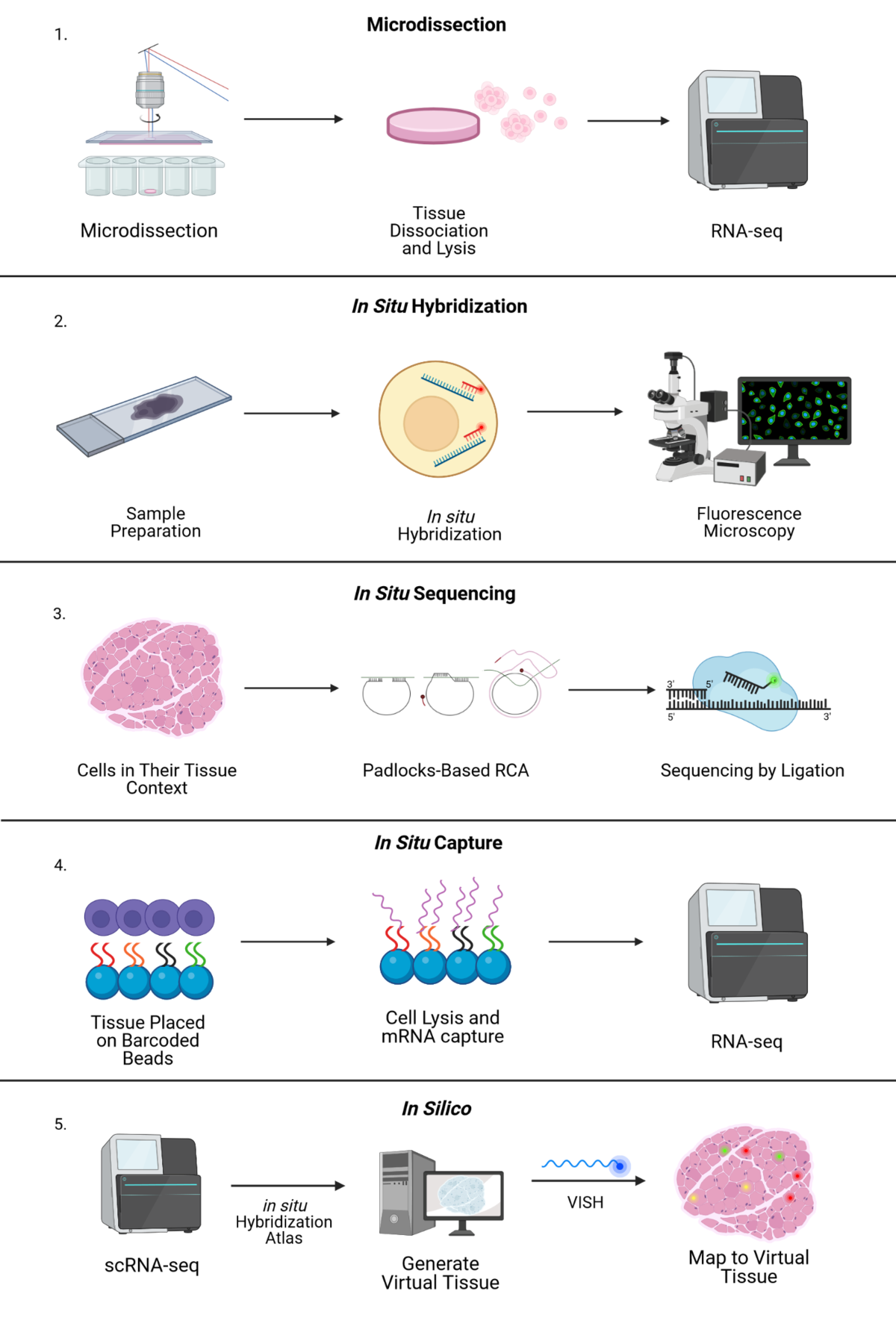
Uncovering an Organ's Molecular Architecture at Single-Cell Resolution by Spatially Resolved Transcriptomics: Trends in Biotechnology
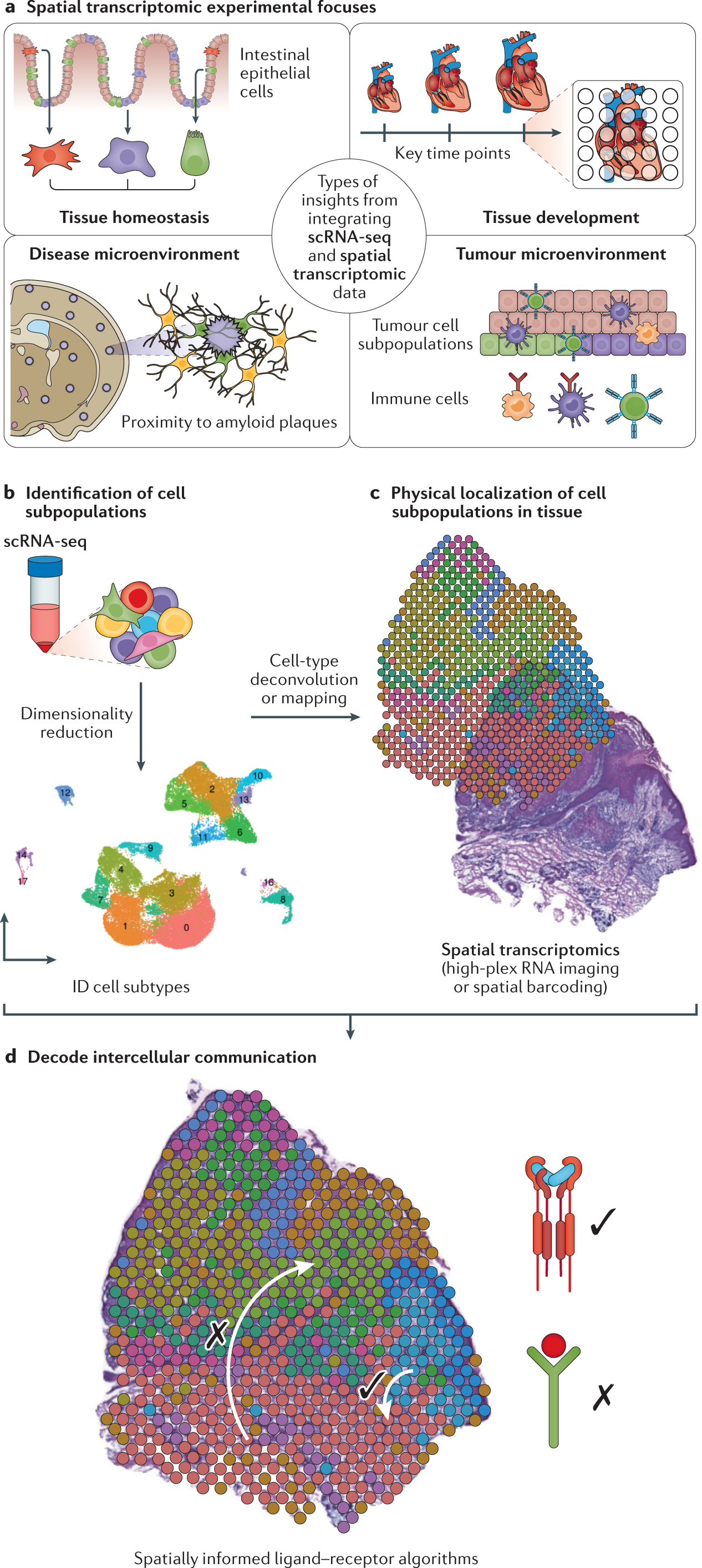
Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics | Nature Reviews Genetics
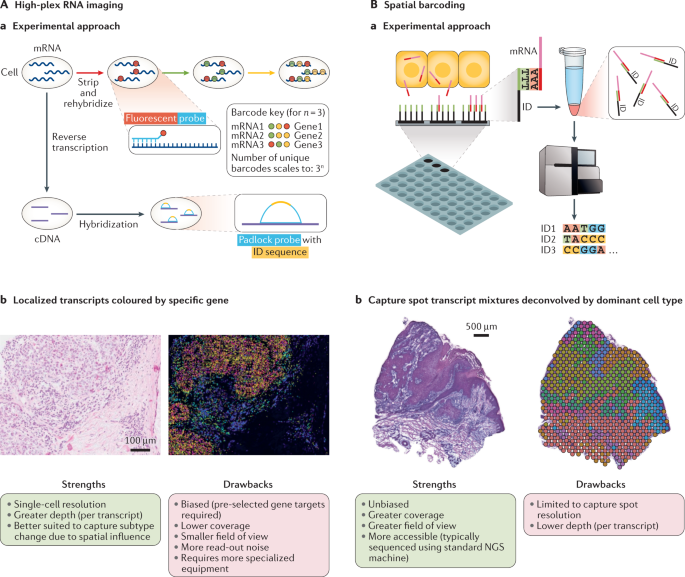
Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics | Nature Reviews Genetics
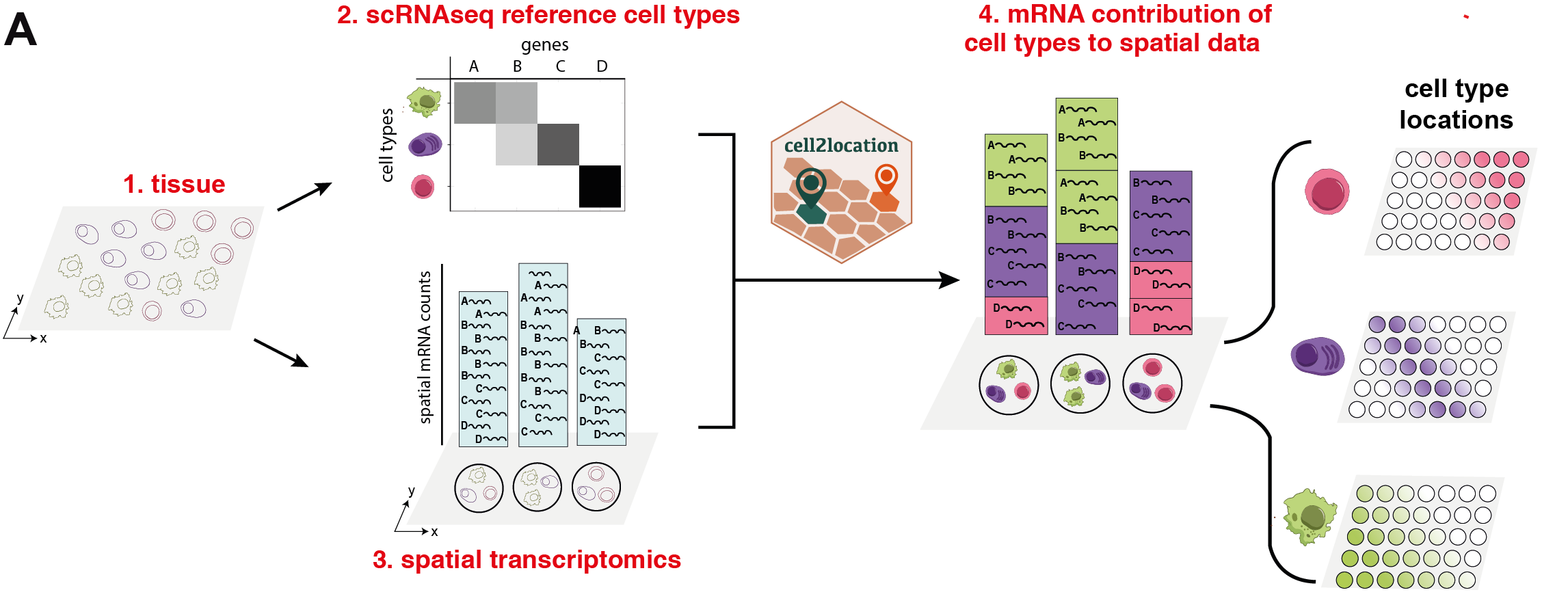
Spatial mapping of cell types across the mouse brain (3/3) - visualising results and downstream analysis — cell2location documentation

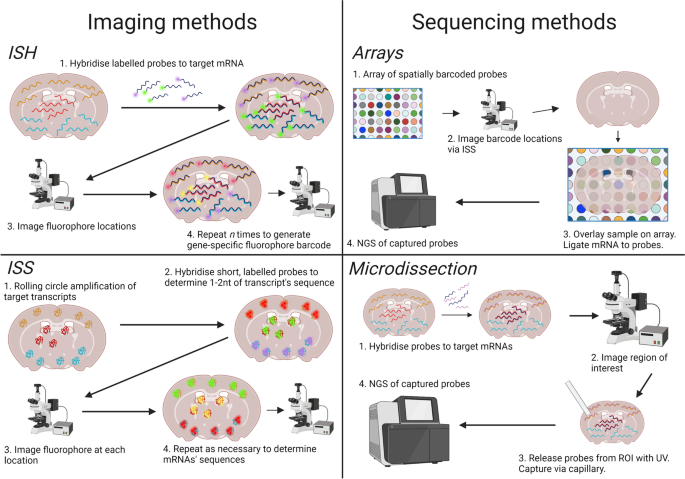
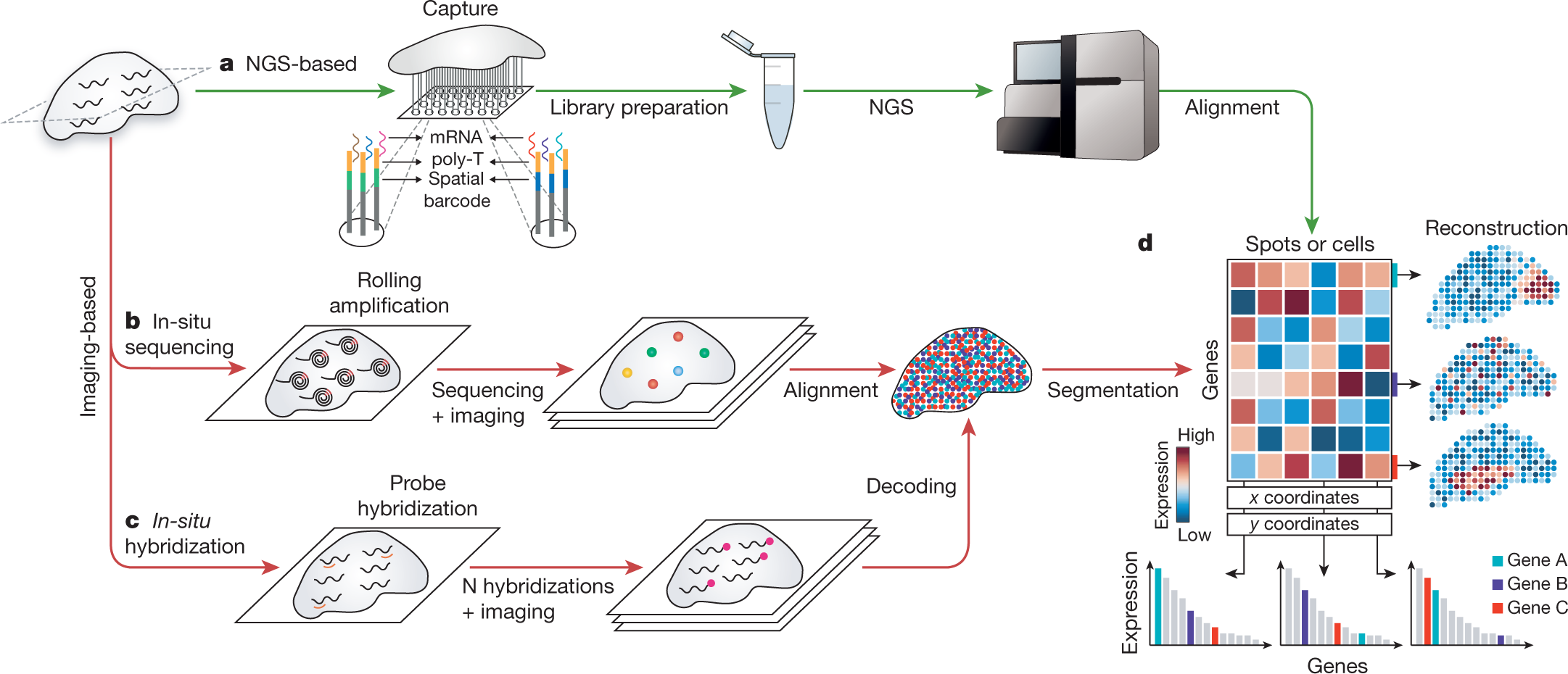
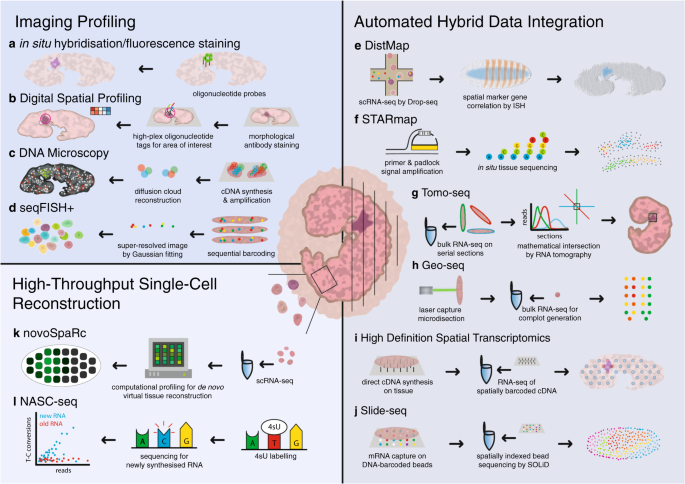
Principles of current methods for capturing spatial gene expression.... | Download Scientific Diagram

Deep Learning in Spatial Transcriptomics: Learning From the Next Next-Generation Sequencing | bioRxiv
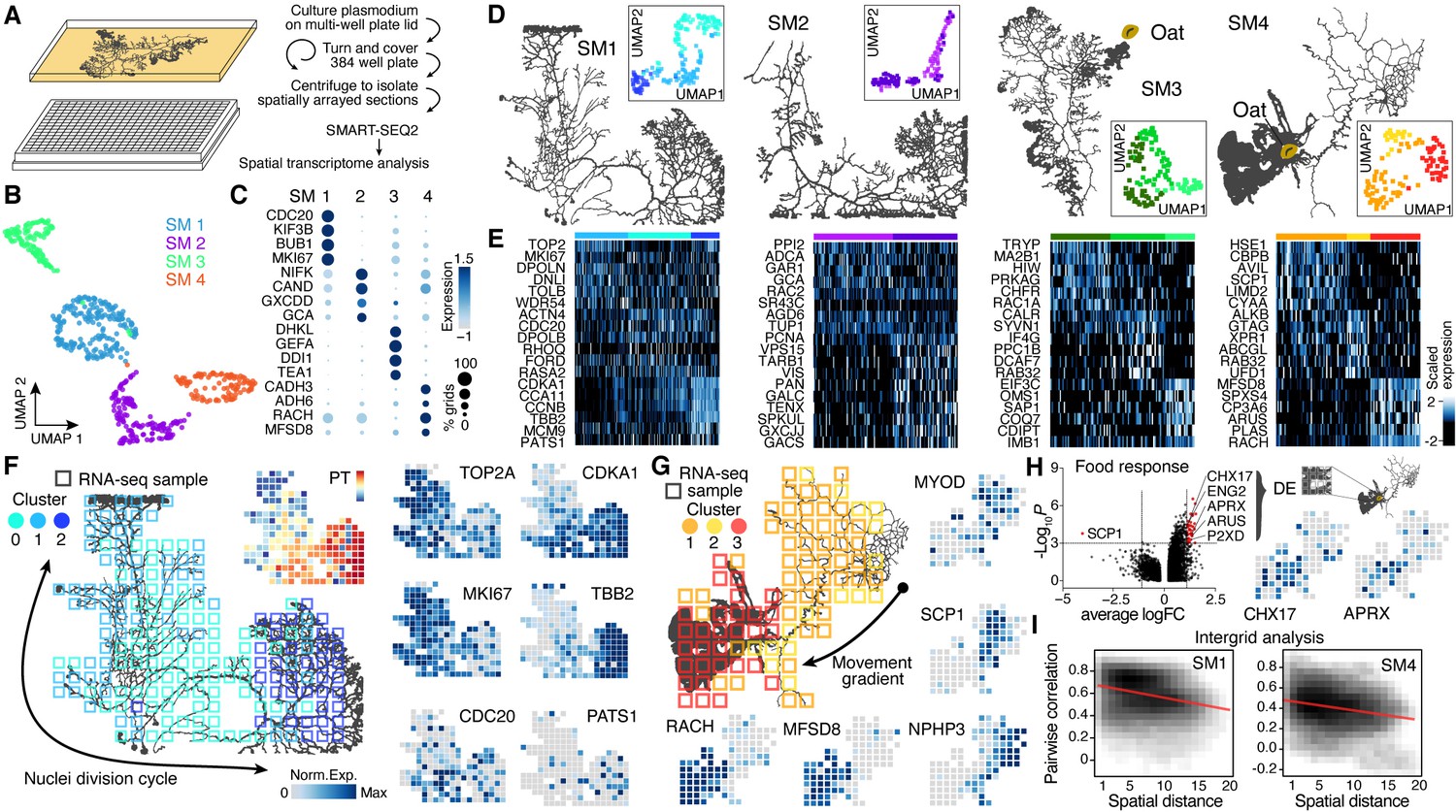
Spatial transcriptomic and single-nucleus analysis reveals heterogeneity in a gigantic single-celled syncytium | eLife

Haiqi Chen on Twitter: "In this work, we built a comprehensive spatial transcriptome atlas for mammalian spermatogenesis at a near single-cell resolution using a high throughput spatial RNA capture technology called Slide-seq. (
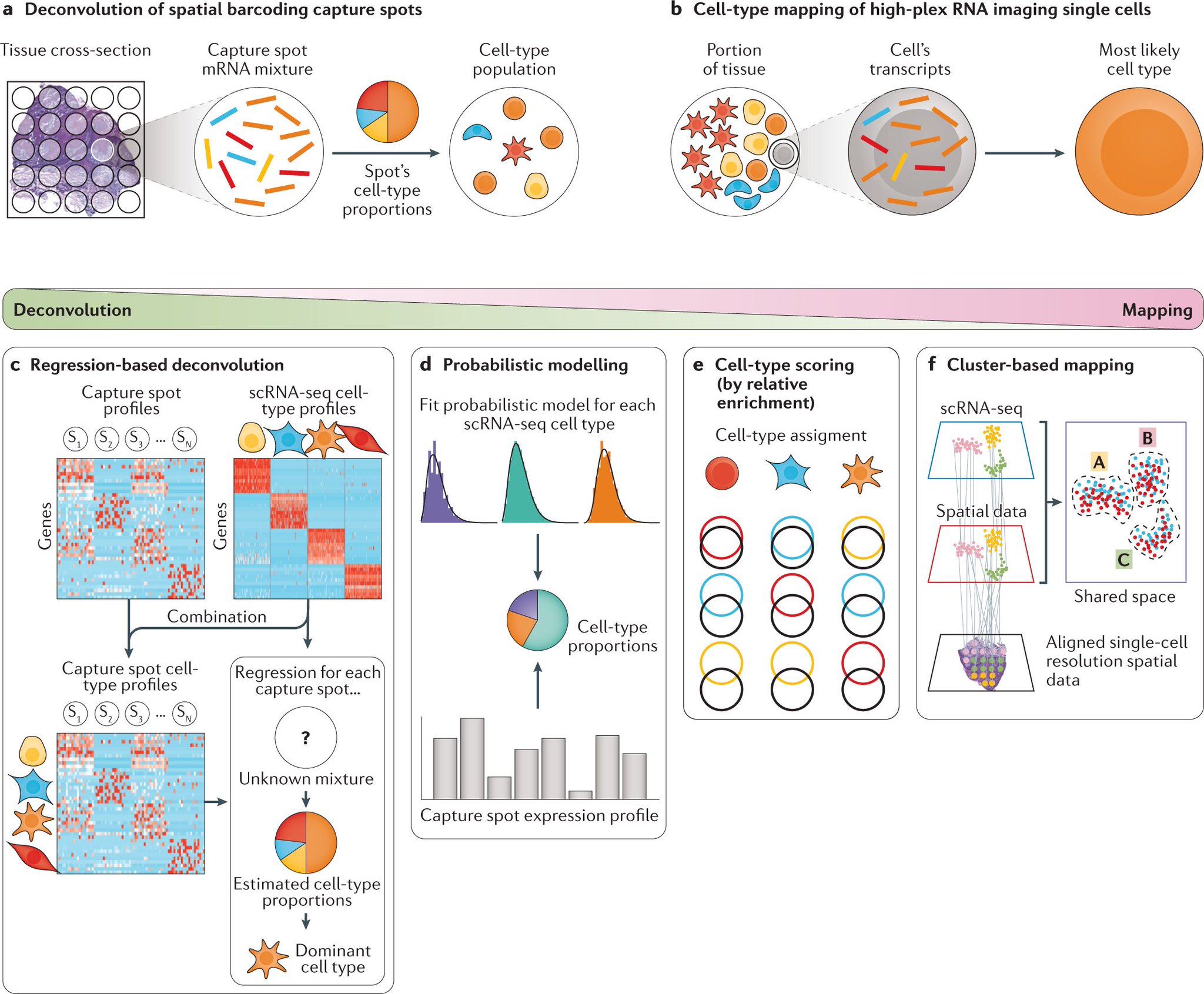
Nature Reviews Genetics on Twitter: "scRNA-seq data can be leveraged to 'deconvolve' cell types at spatial barcoding capture spots and to 'map' subtypes to high-plex RNA imaging (includes multiplexed in situ hybridization
From whole-mount to single-cell spatial assessment of gene expression in 3D | Communications Biology