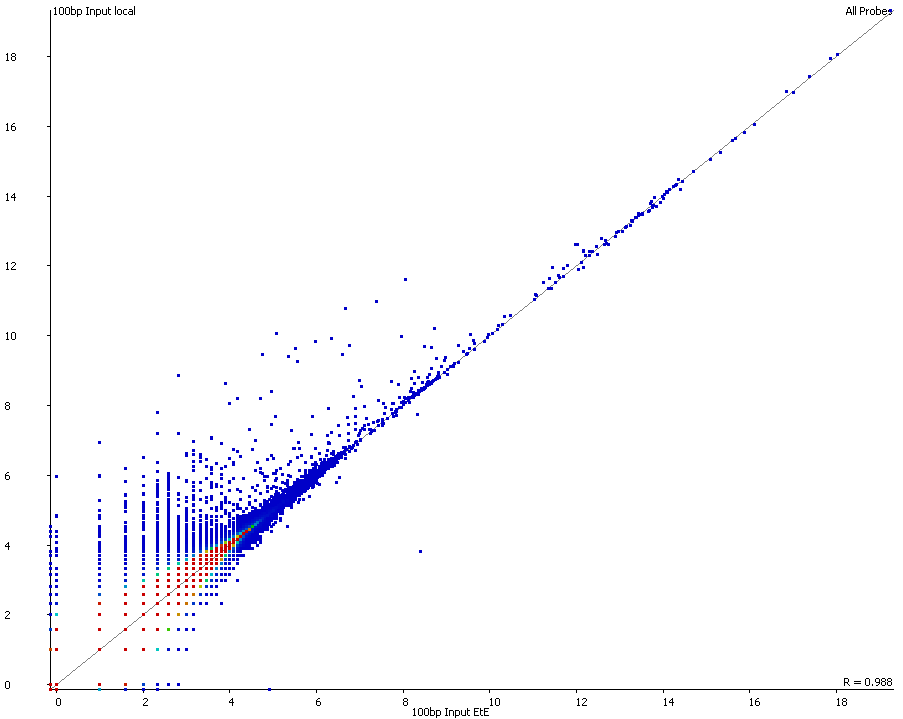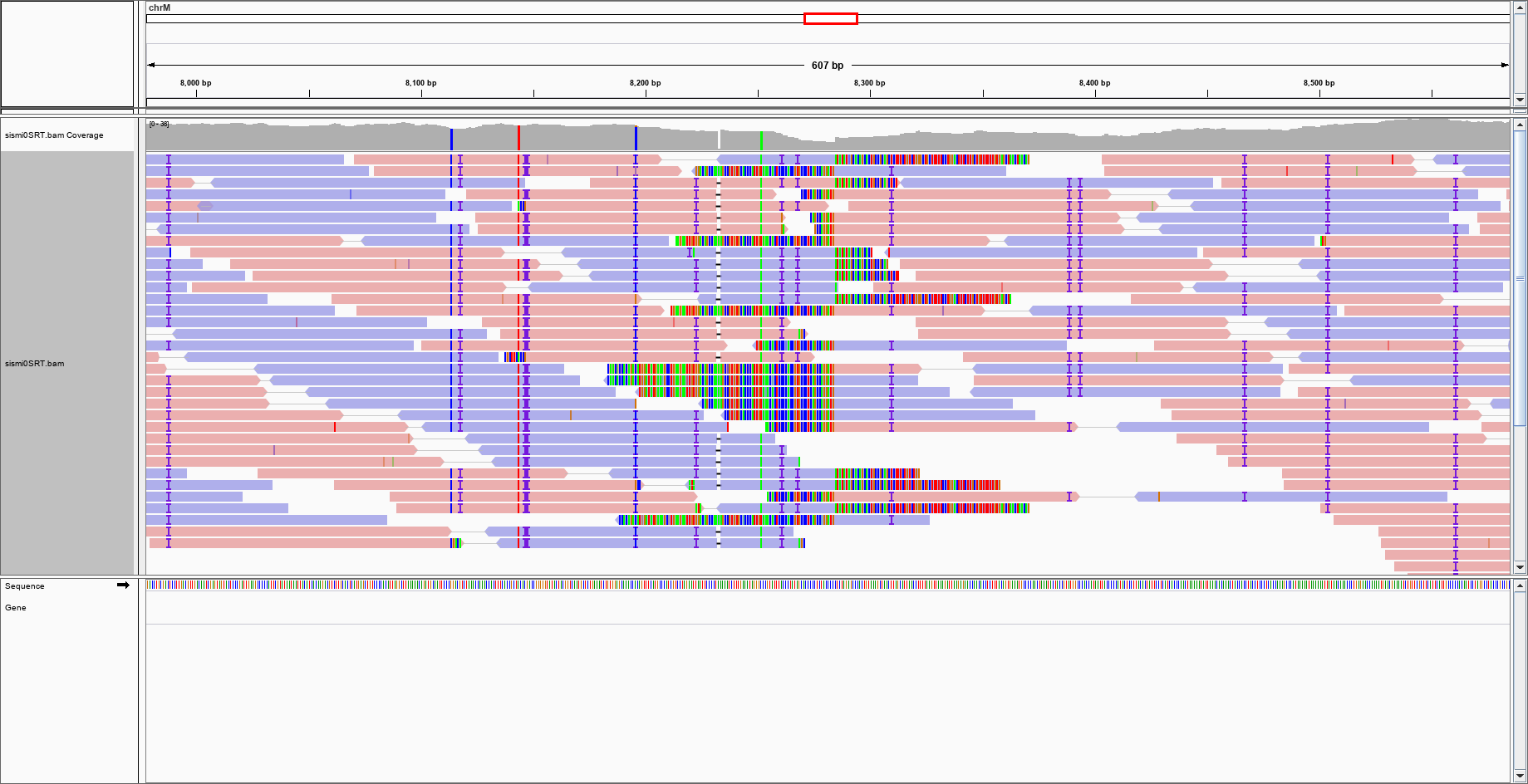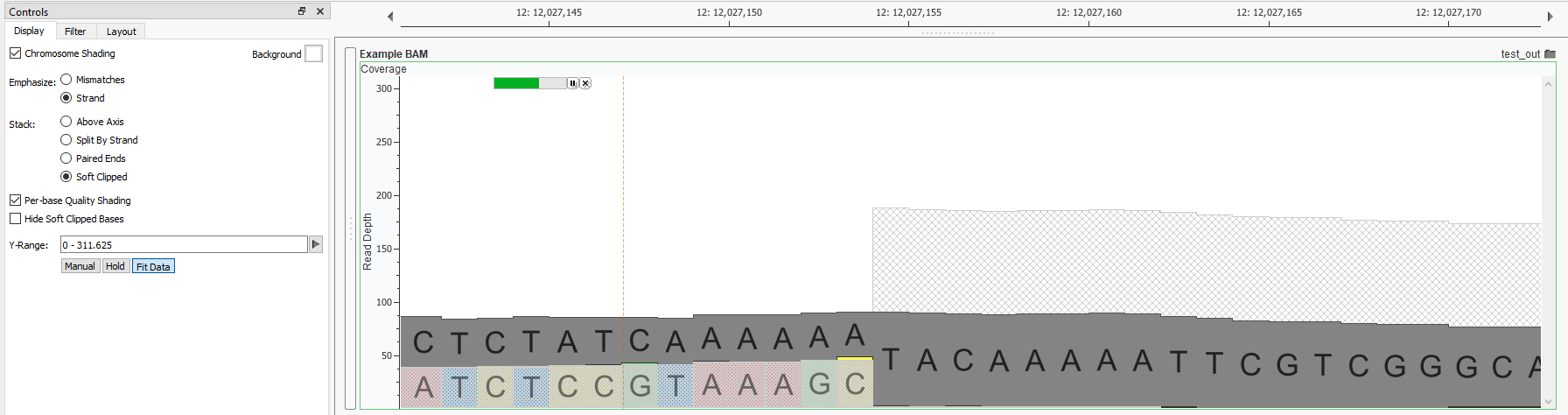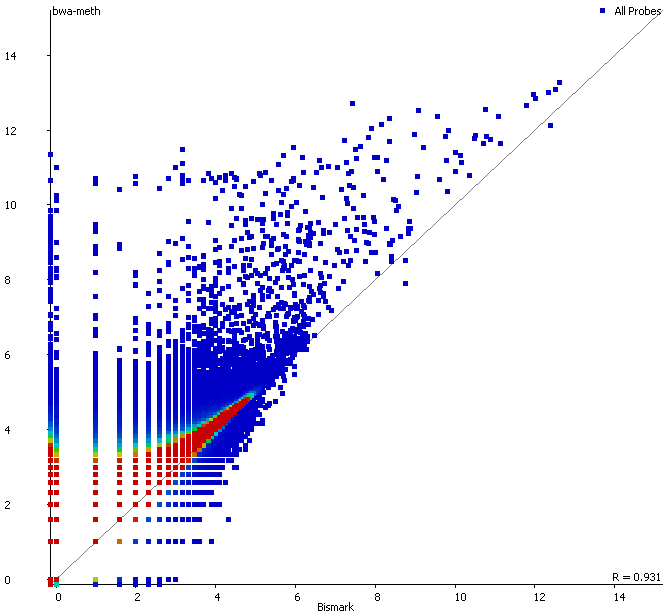
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions

Strange differences in detected variants called by mutect2 --dont-use-soft- clipped-bases true vs false – GATK

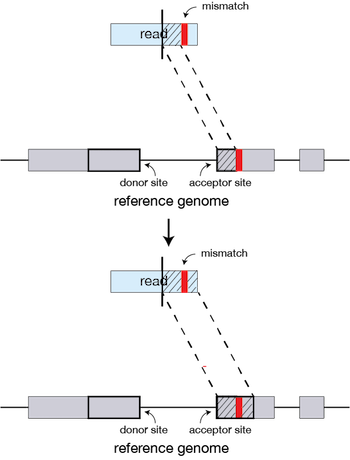
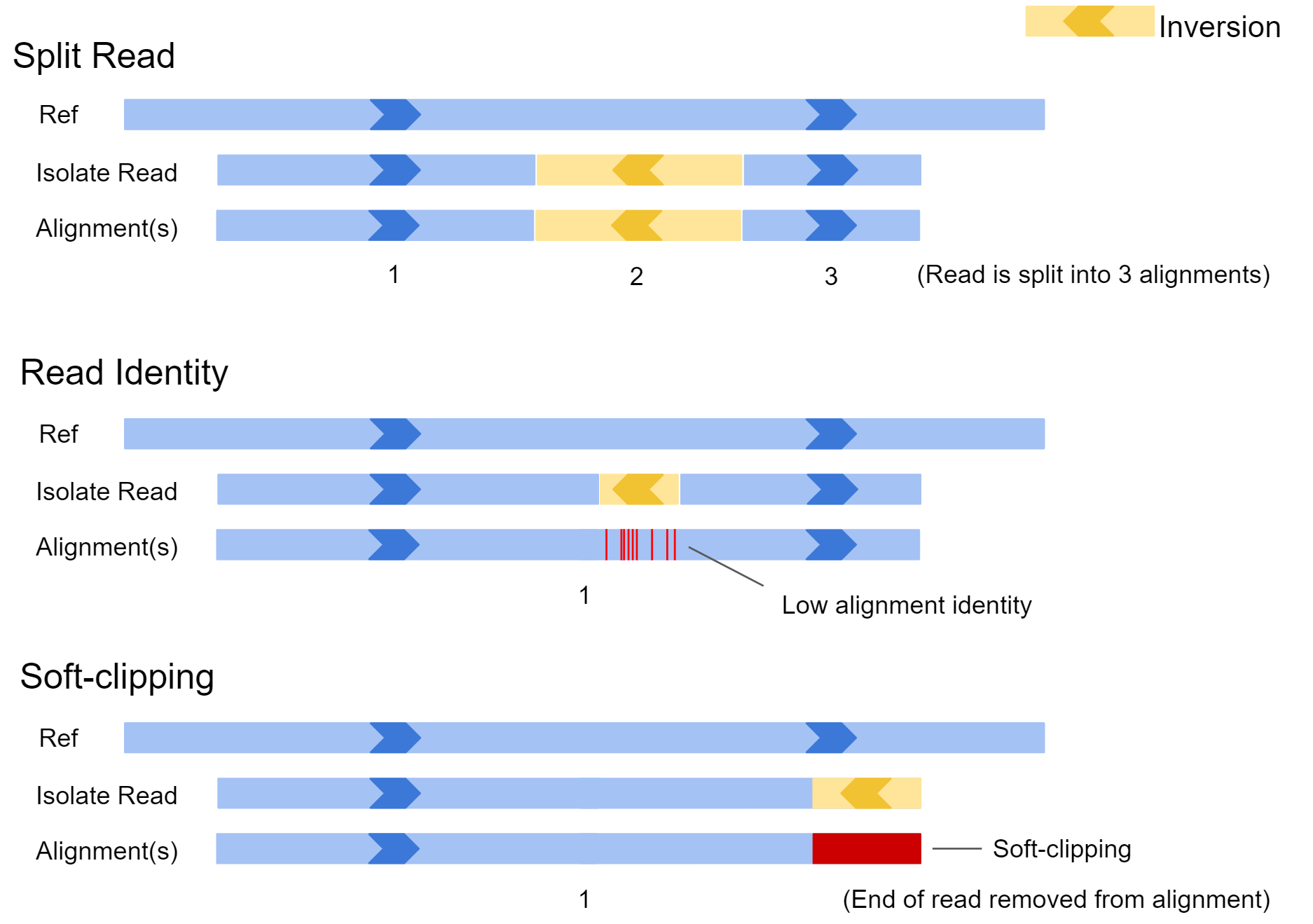
Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram

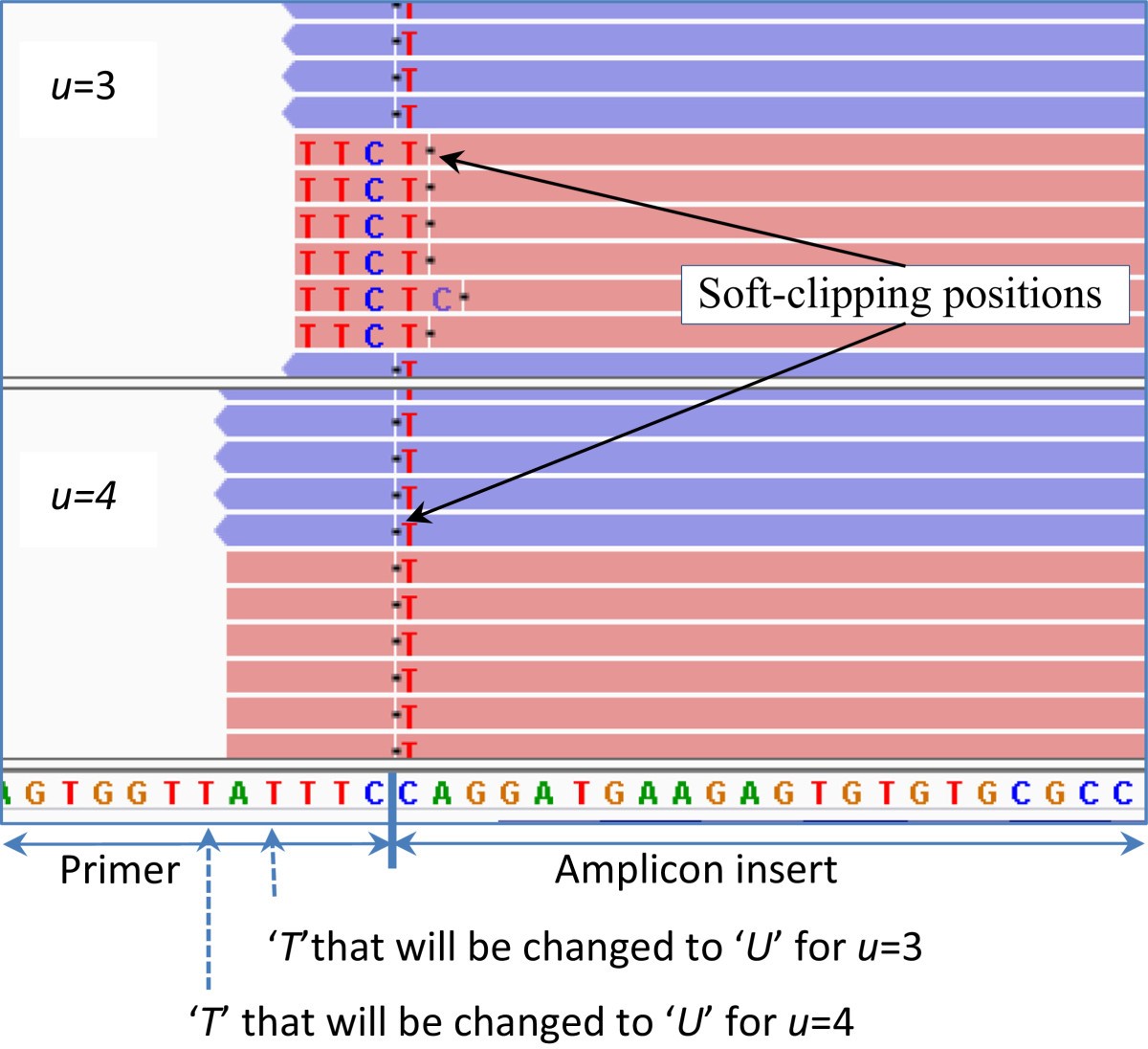
Common sources of 5′end soft clipping and mismatches in 3′ end seq. (a)... | Download Scientific Diagram

Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram

Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram
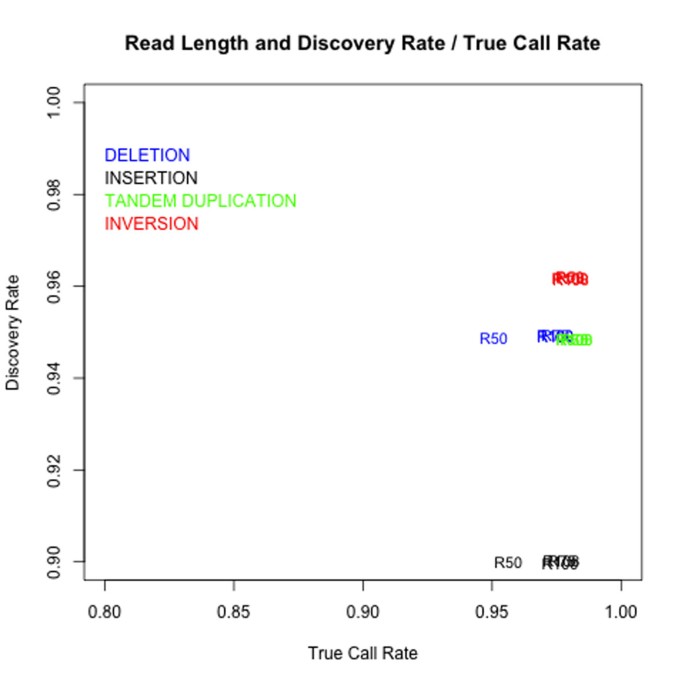
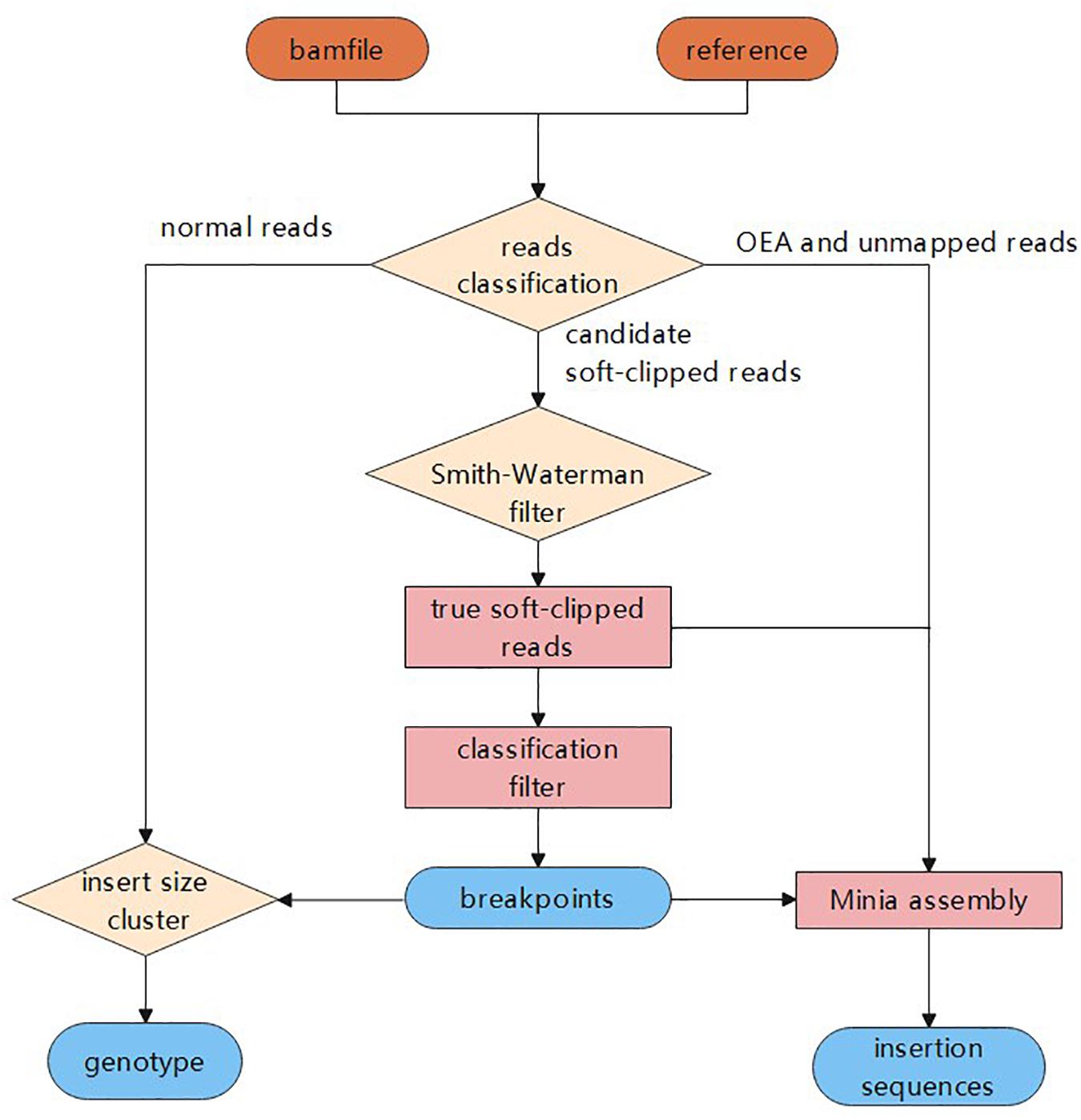
ClipCrop: a tool for detecting structural variations with single-base resolution using soft-clipping information | BMC Bioinformatics | Full Text

IMSindel: An accurate intermediate-size indel detection tool incorporating de novo assembly and gapped global-local alignment with split read analysis | Scientific Reports
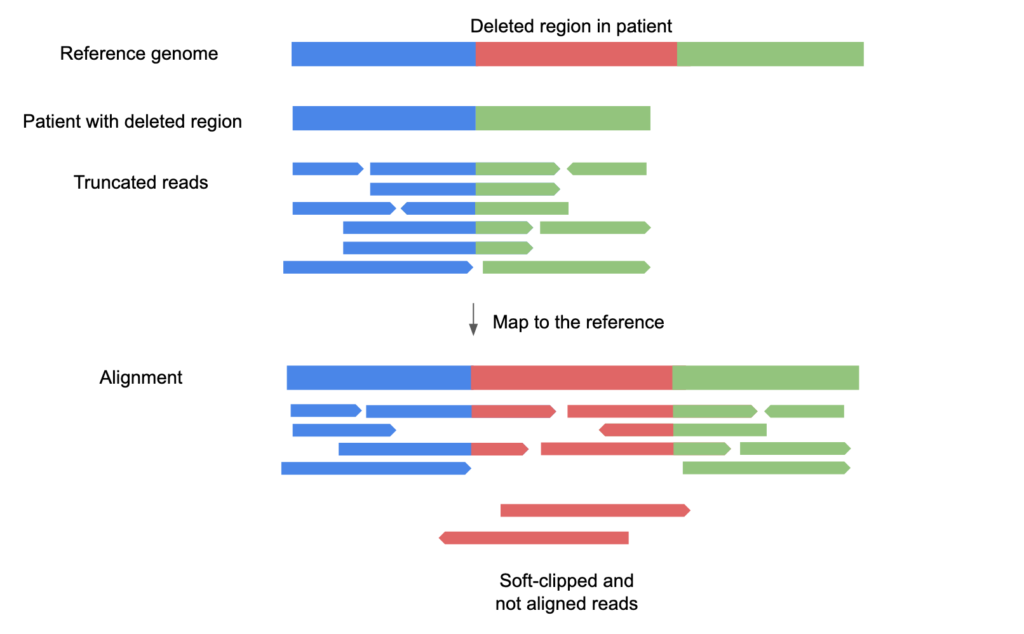
Challenges and importance of mid-sized deletion identification for genomic medicine - SeqOne Genomics
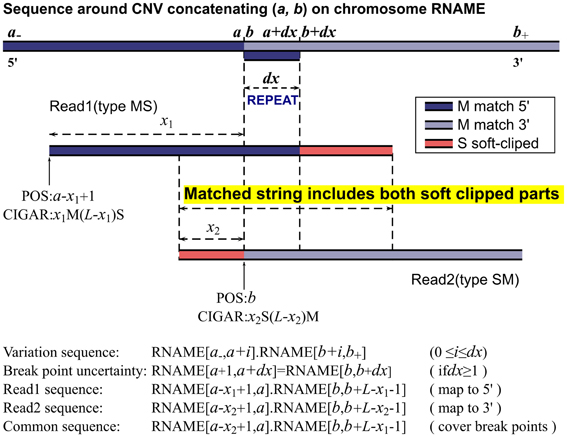
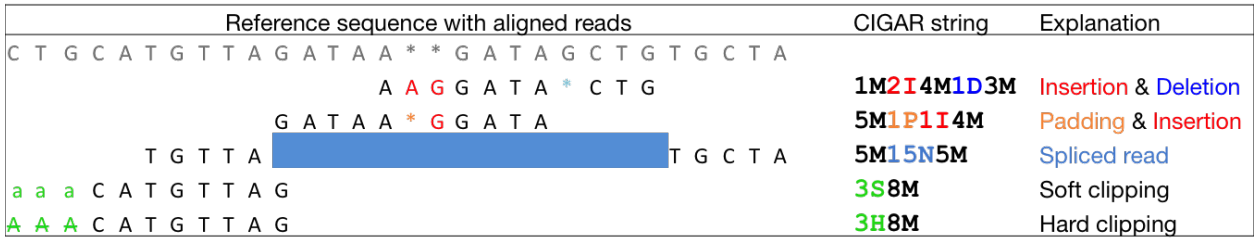
Frontiers | MATCHCLIP: locate precise breakpoints for copy number variation using CIGAR string by matching soft clipped reads
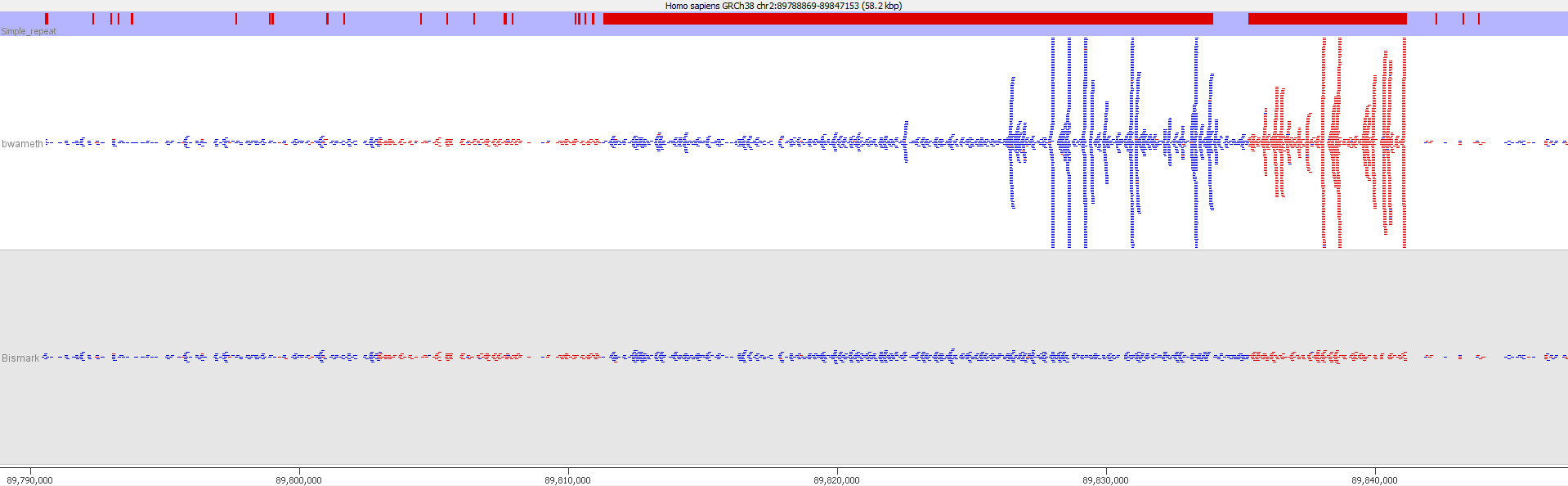
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions
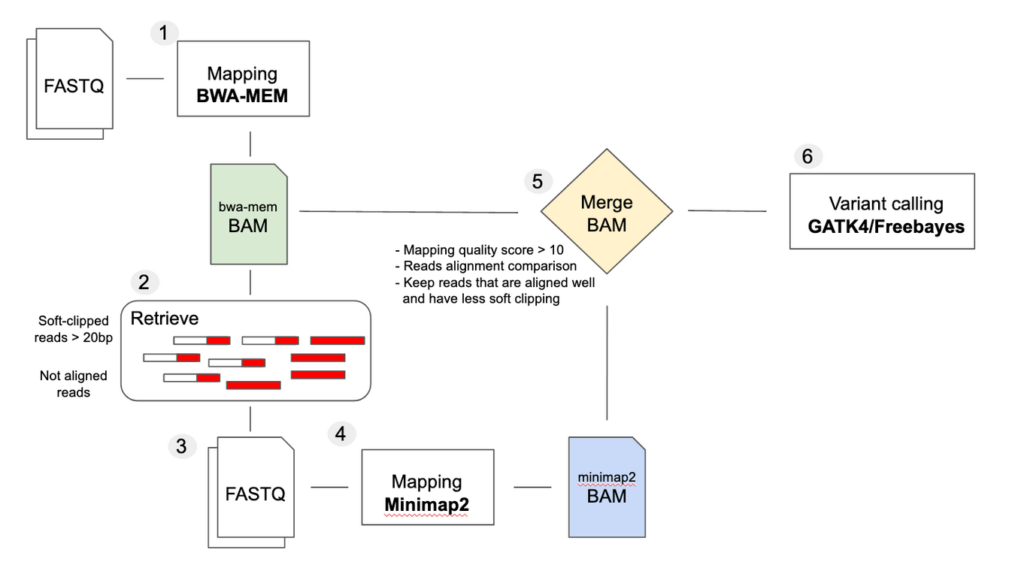
Challenges and importance of mid-sized deletion identification for genomic medicine - SeqOne Genomics

Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram
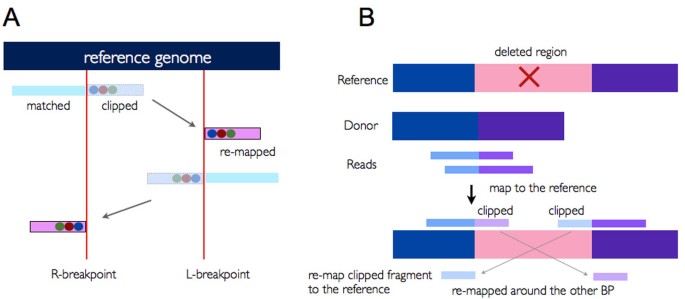
ClipCrop: a tool for detecting structural variations with single-base resolution using soft-clipping information | BMC Bioinformatics | Full Text