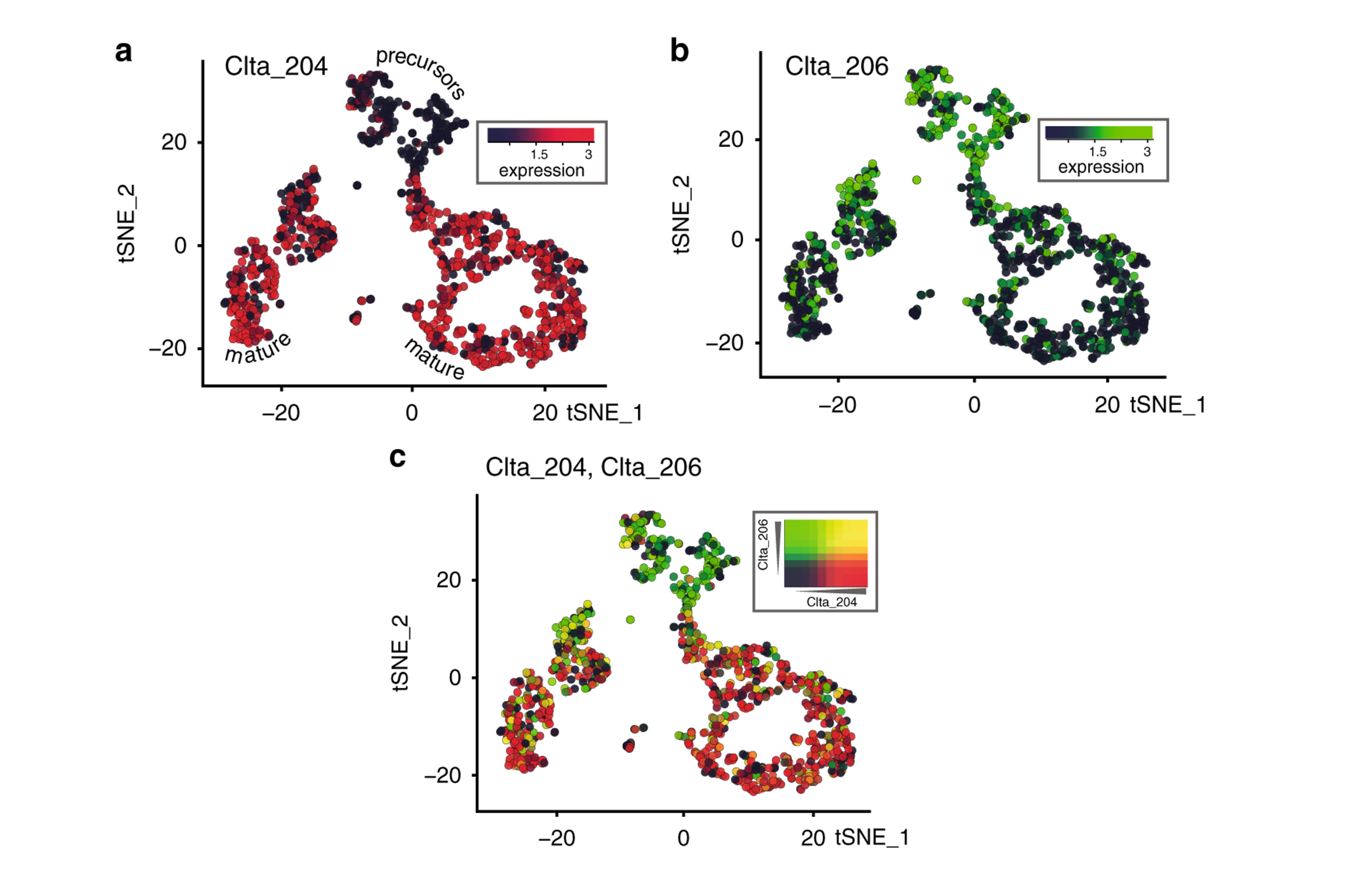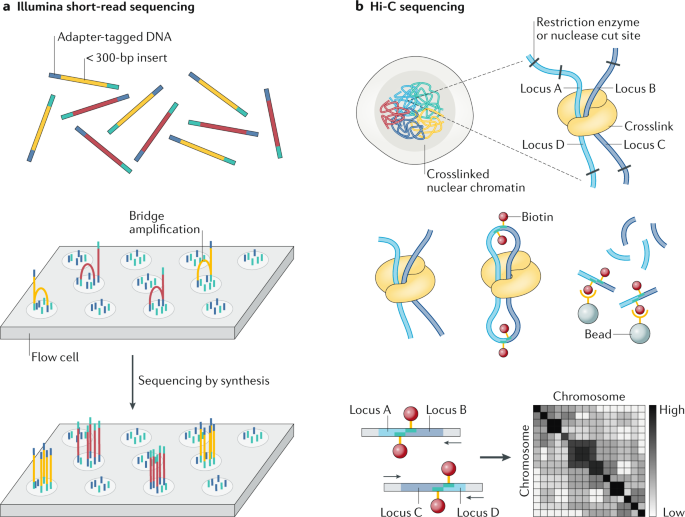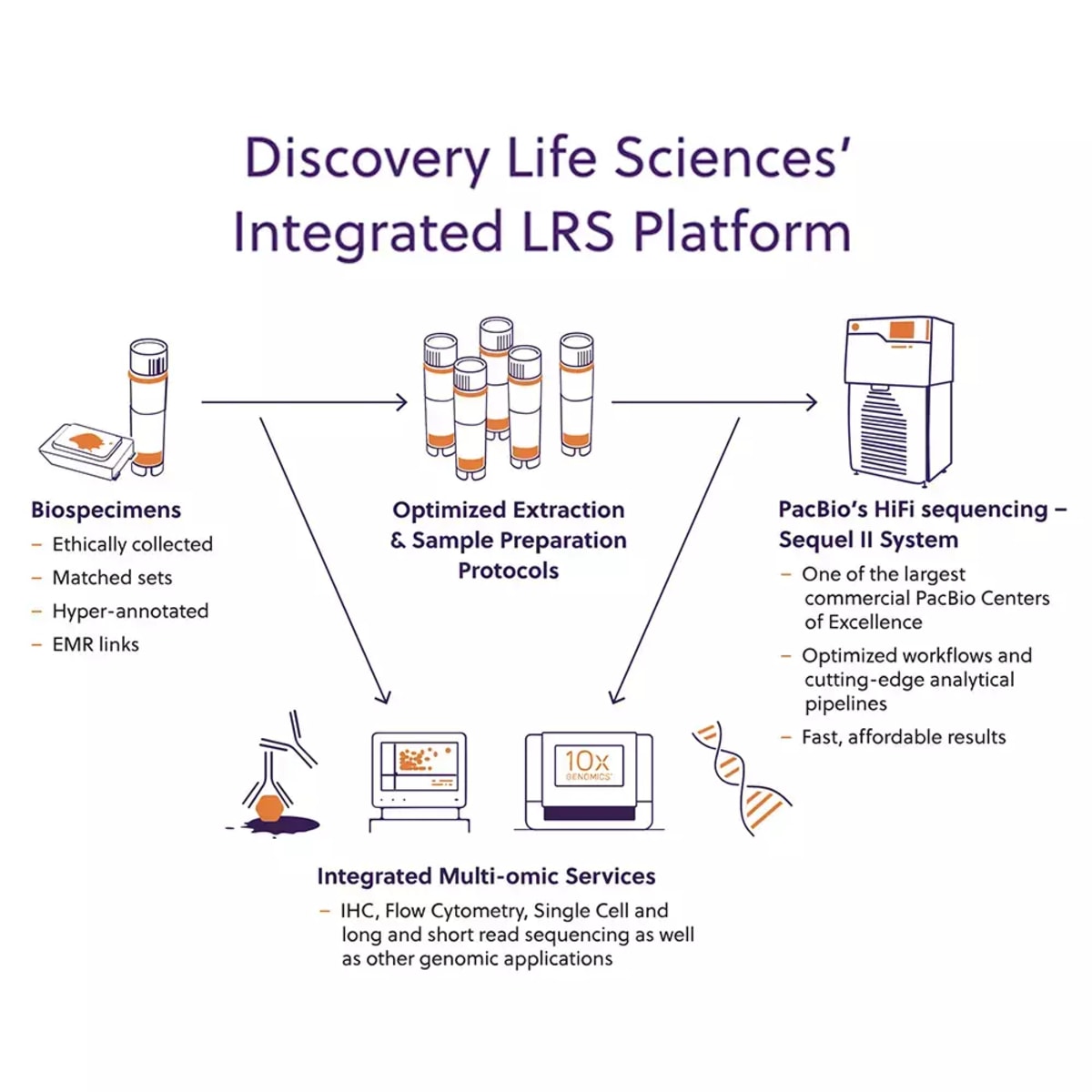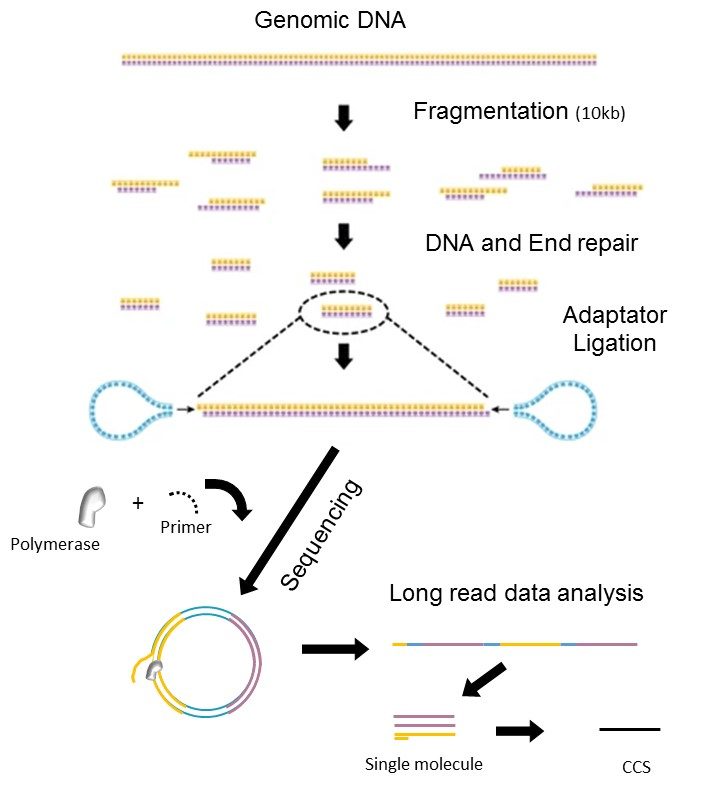
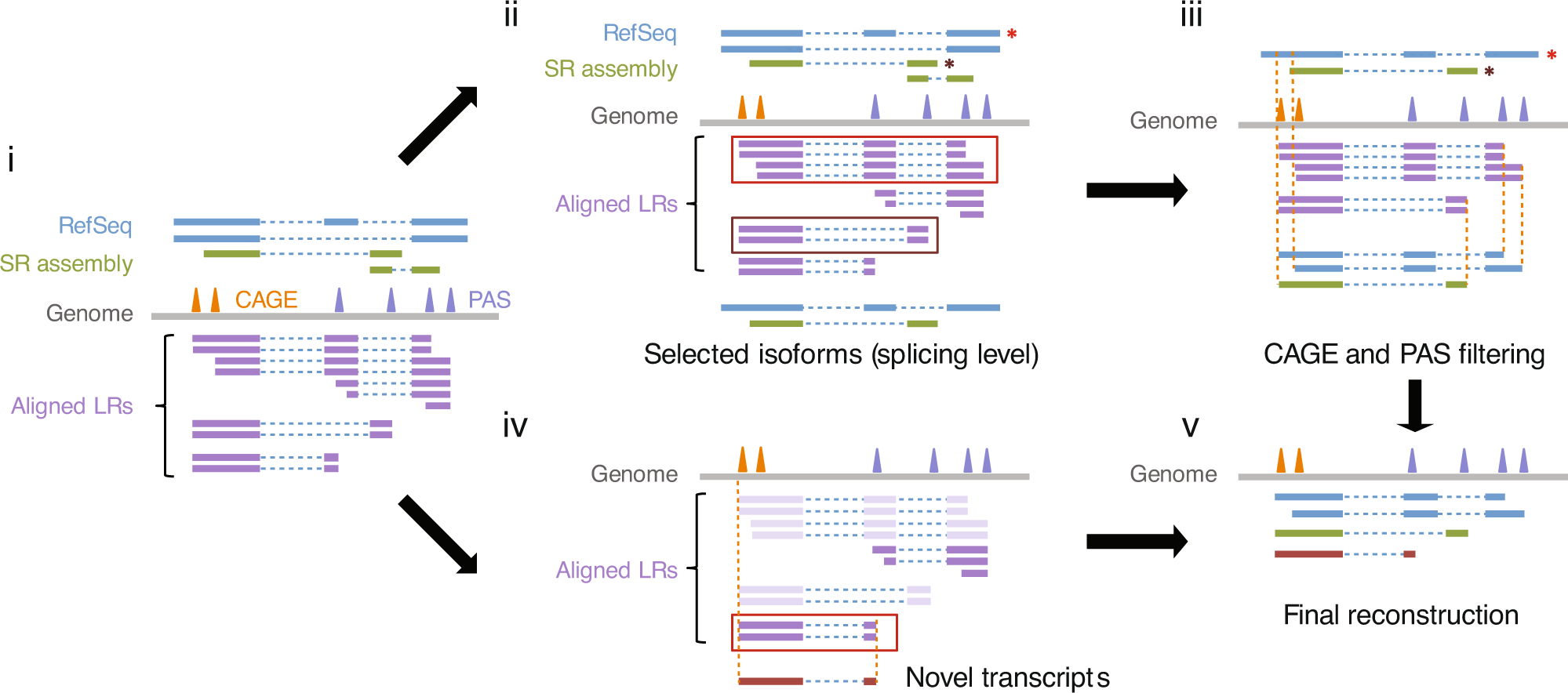
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing | Genome Biology | Full Text

High throughput error corrected Nanopore single cell transcriptome sequencing | Nature Communications
Single-molecule long-read sequencing reveals a conserved intact long RNA profile in sperm | Nature Communications

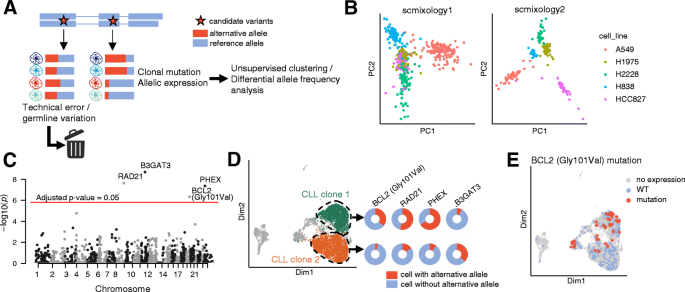
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes | Nature Communications

Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology
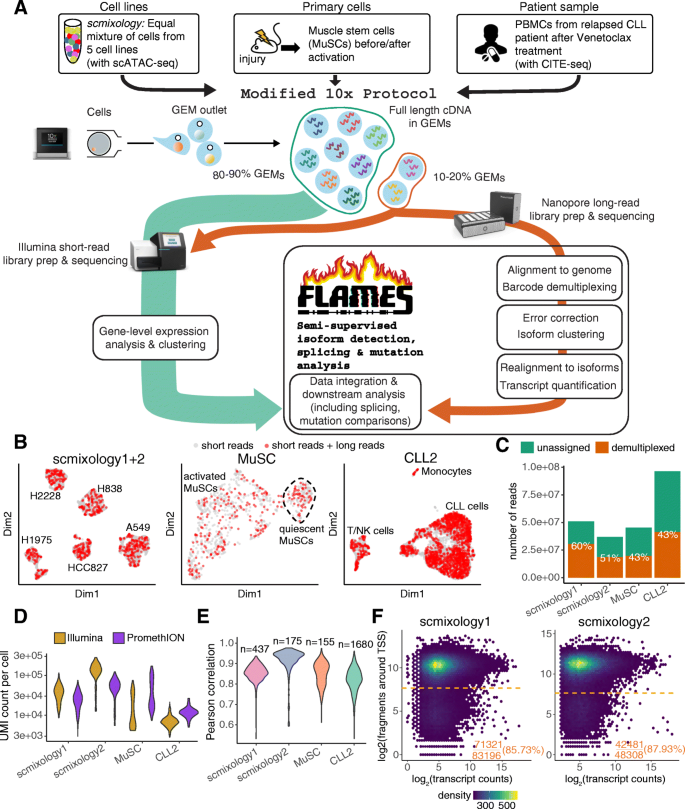
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing | Genome Biology | Full Text
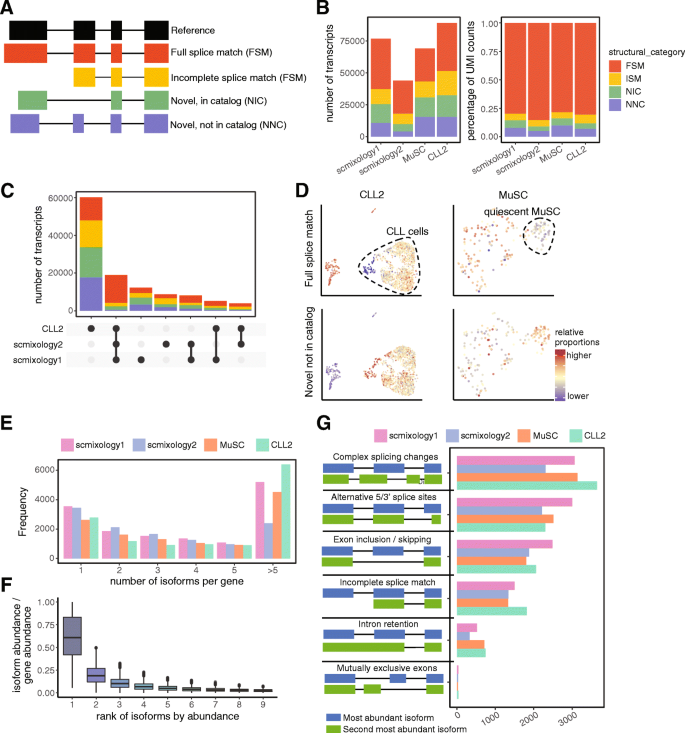
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing | Genome Biology | Full Text
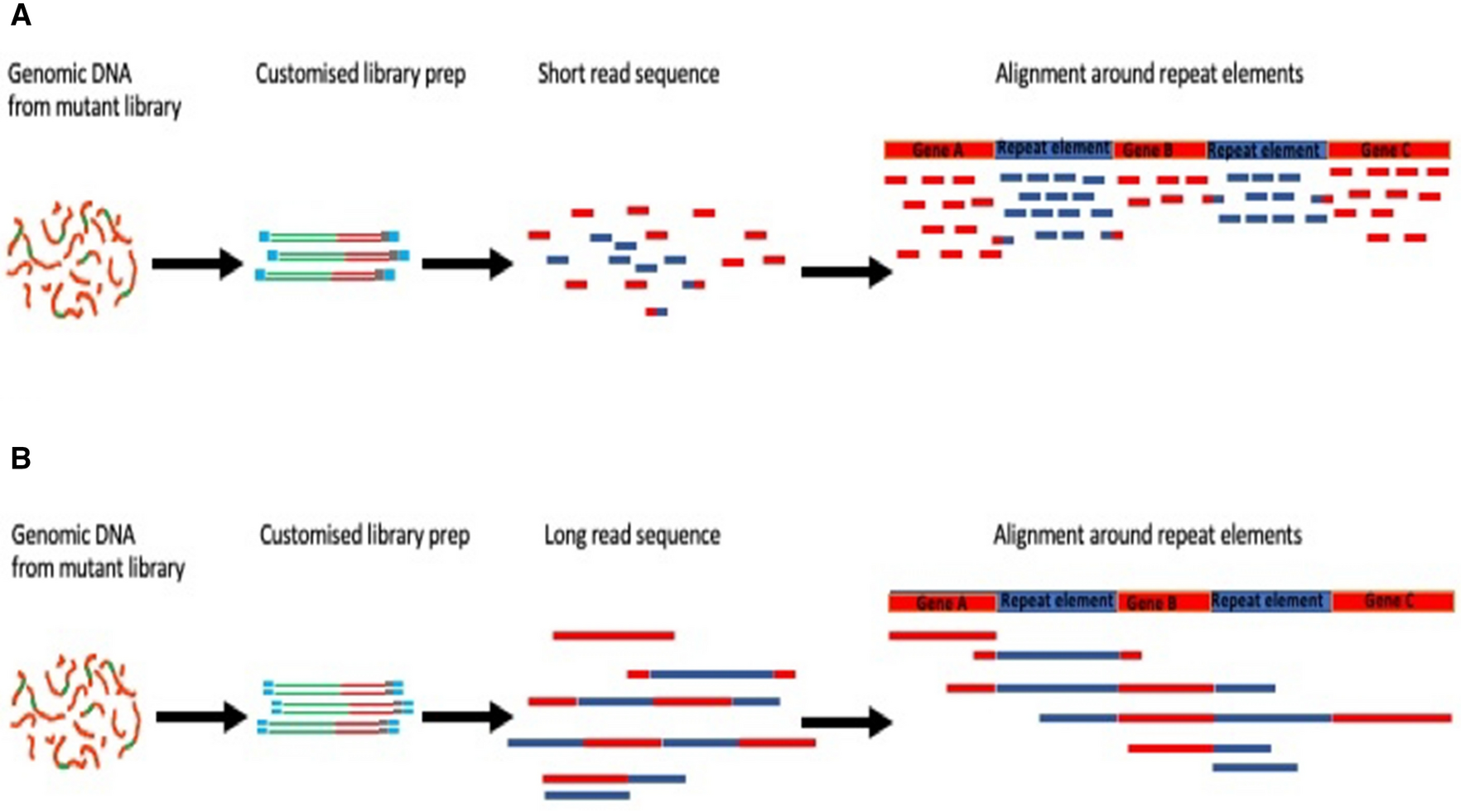
Long-read sequencing for identification of insertion sites in large transposon mutant libraries | Scientific Reports
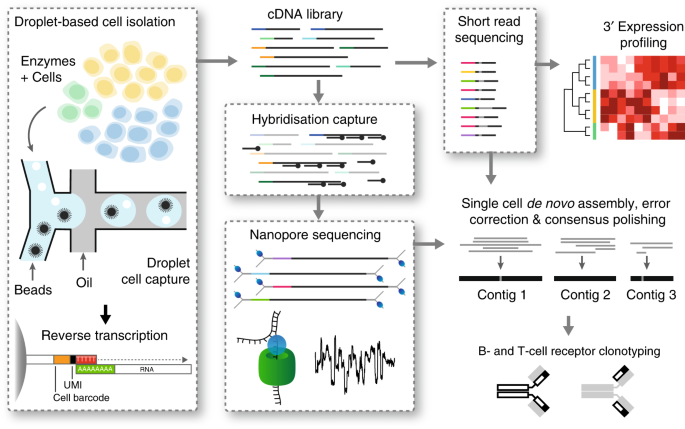
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes | Nature Communications
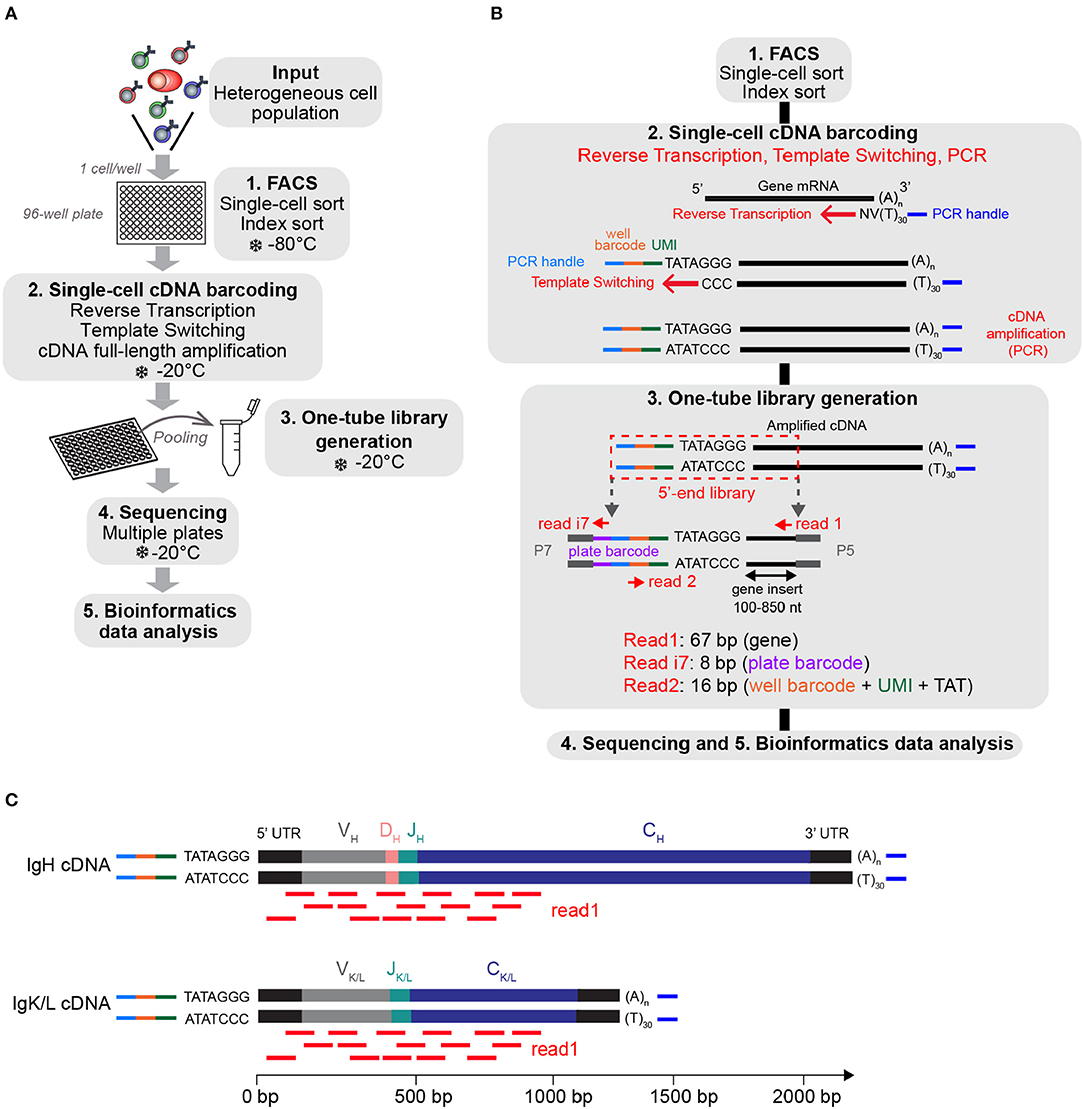
Frontiers | FB5P-seq: FACS-Based 5-Prime End Single-Cell RNA-seq for Integrative Analysis of Transcriptome and Antigen Receptor Repertoire in B and T Cells