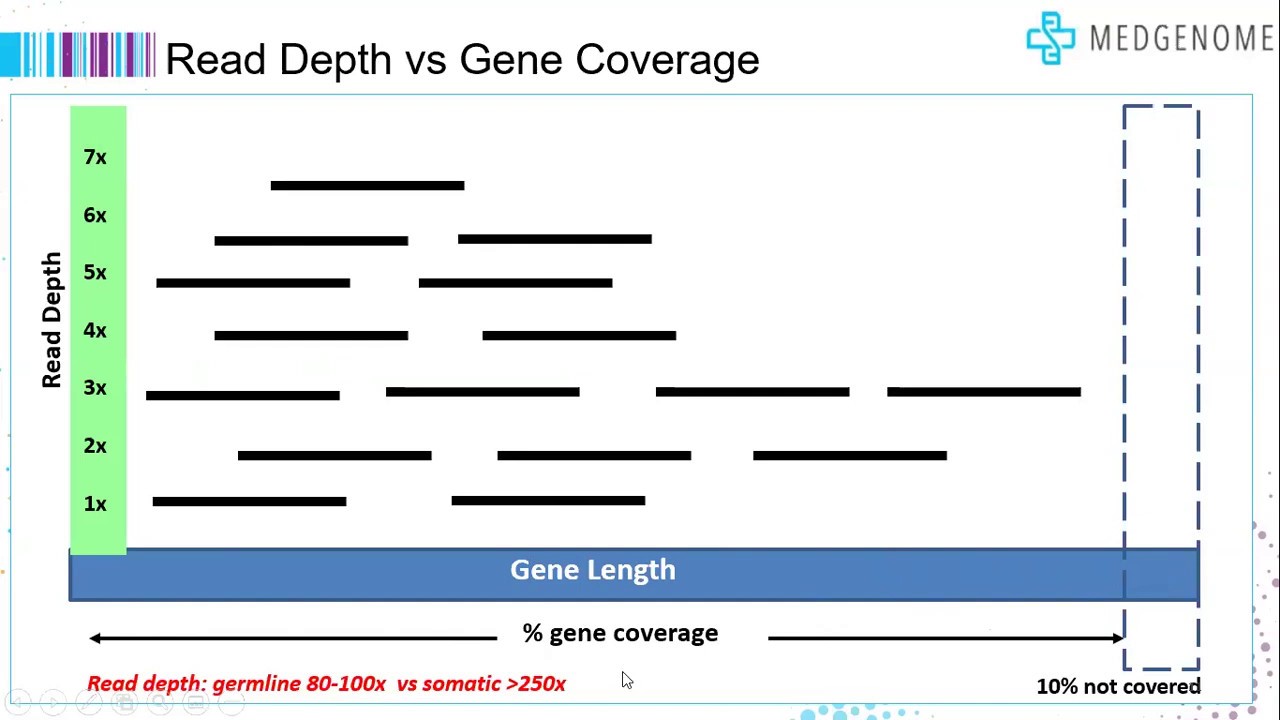
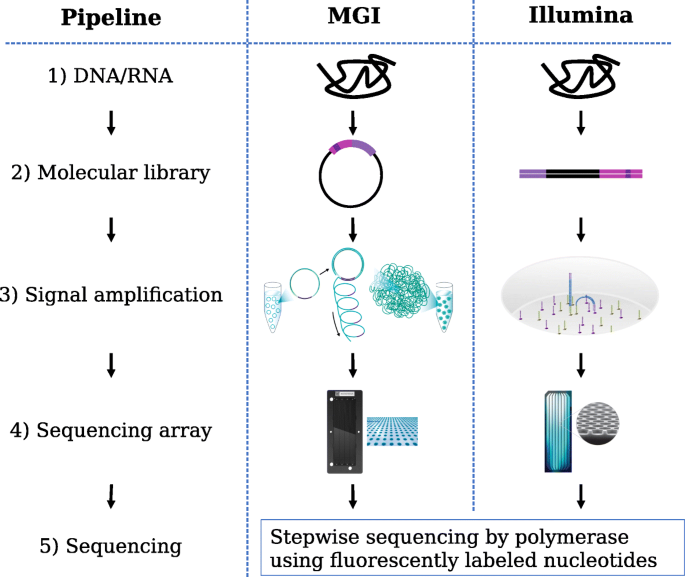
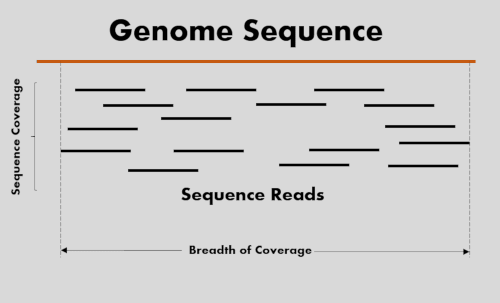
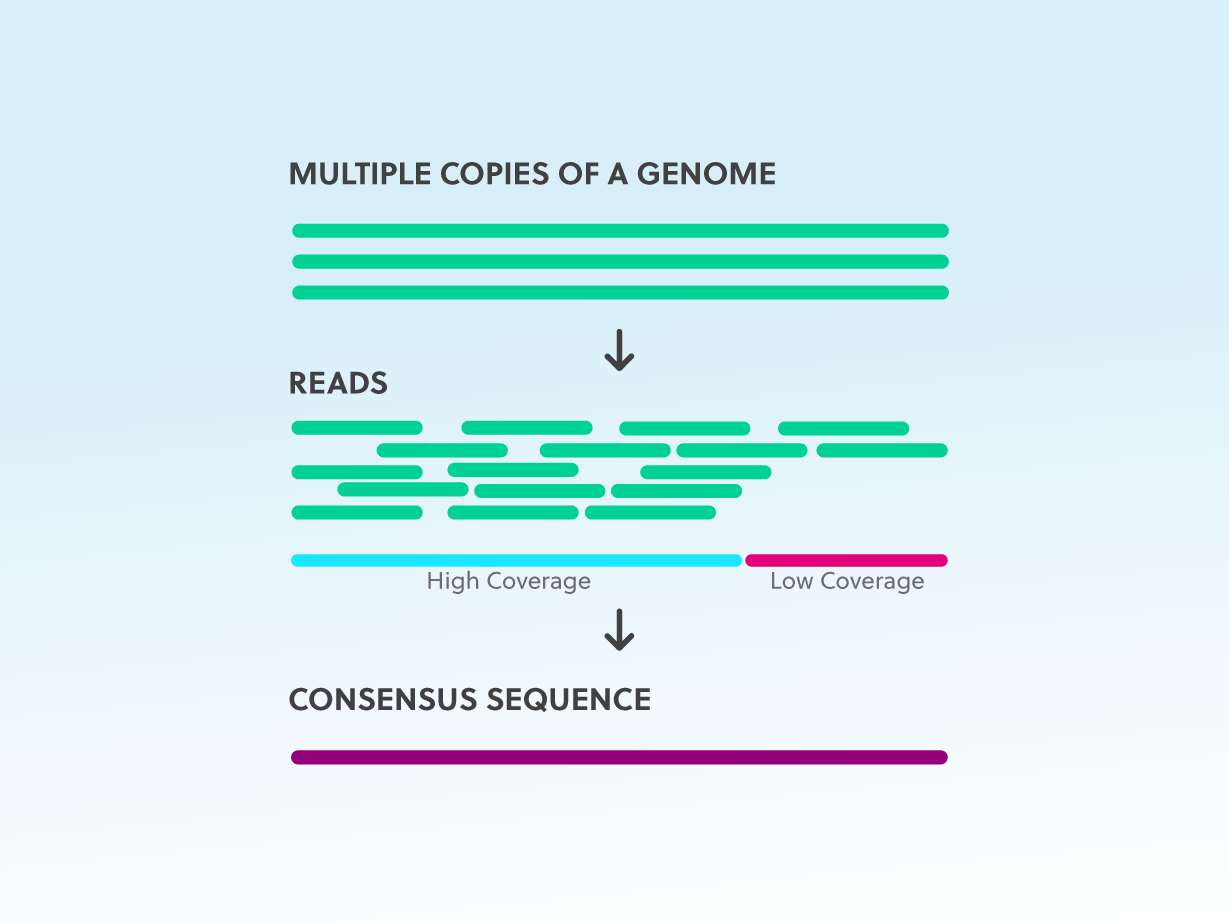
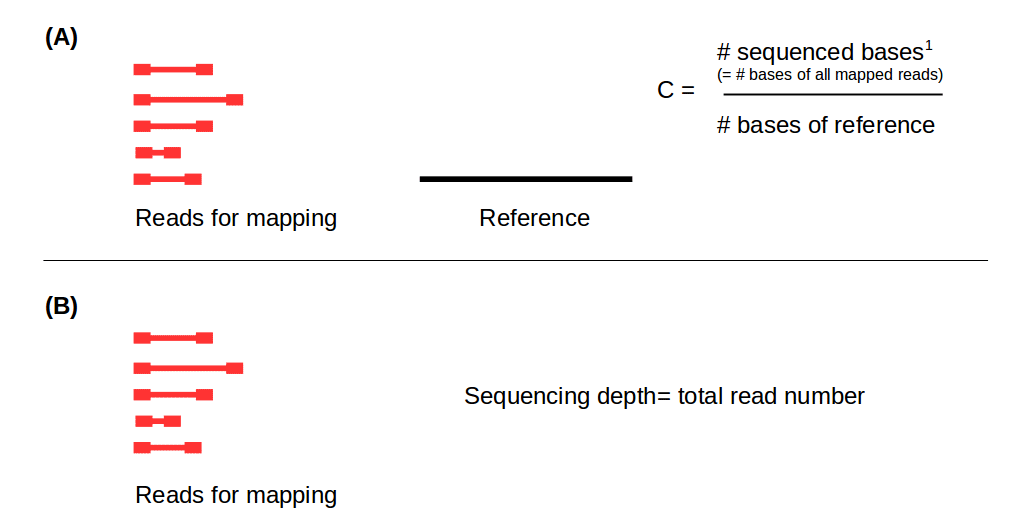
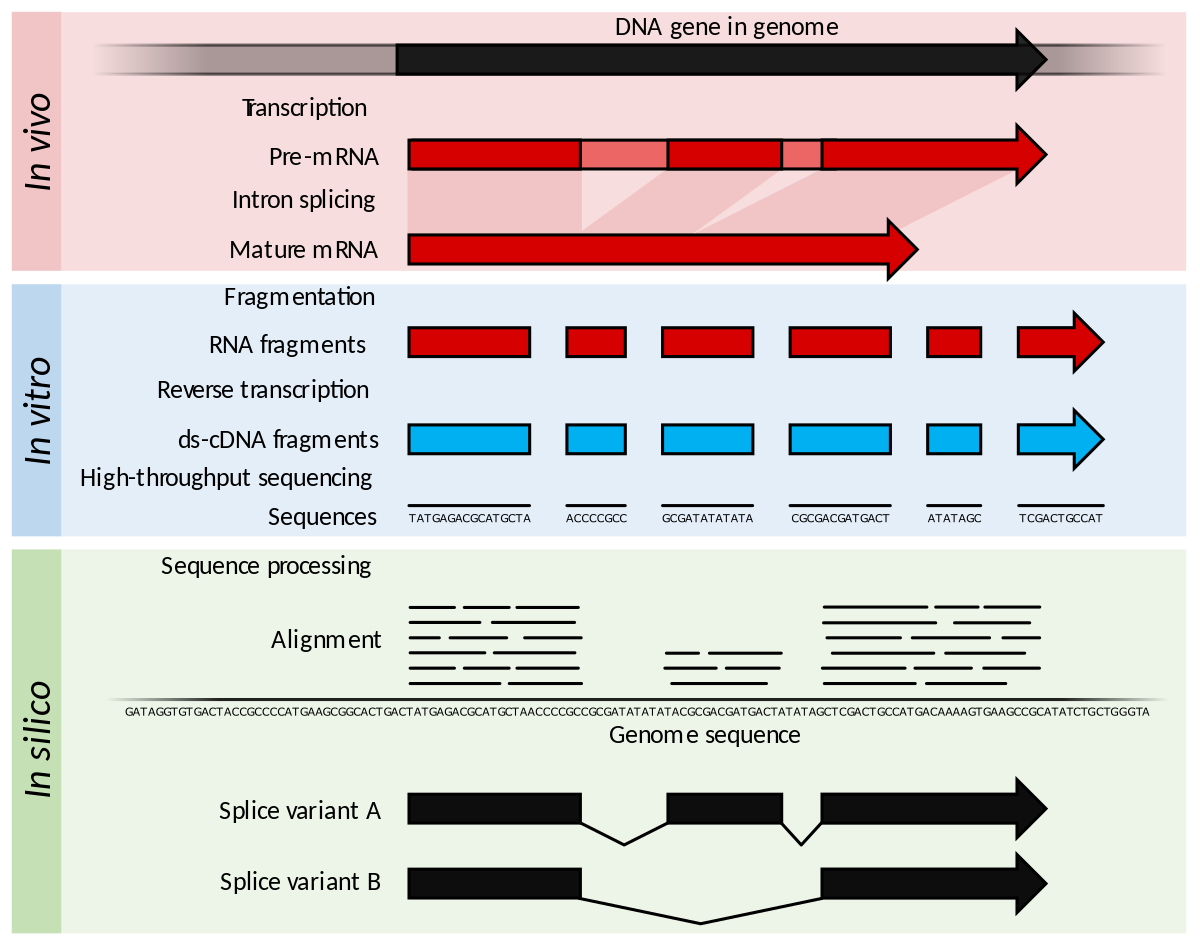
Read coverage of genome sequencing and RNA-seq data. a Read coverage of... | Download Scientific Diagram

Detecting, Categorizing, and Correcting Coverage Anomalies of RNA-Seq Quantification - ScienceDirect
Improved Annotation of 3′ Untranslated Regions and Complex Loci by Combination of Strand-Specific Direct RNA Sequencing, RNA-Seq and ESTs | PLOS ONE
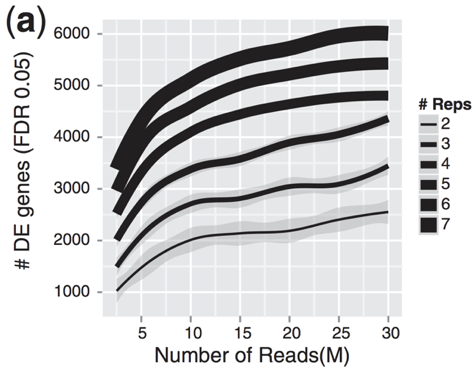
Experimental design considerations | Introduction to RNA-Seq using high-performance computing - ARCHIVED

Determination of the number of reads needed for each RNA-Seq protocol... | Download Scientific Diagram
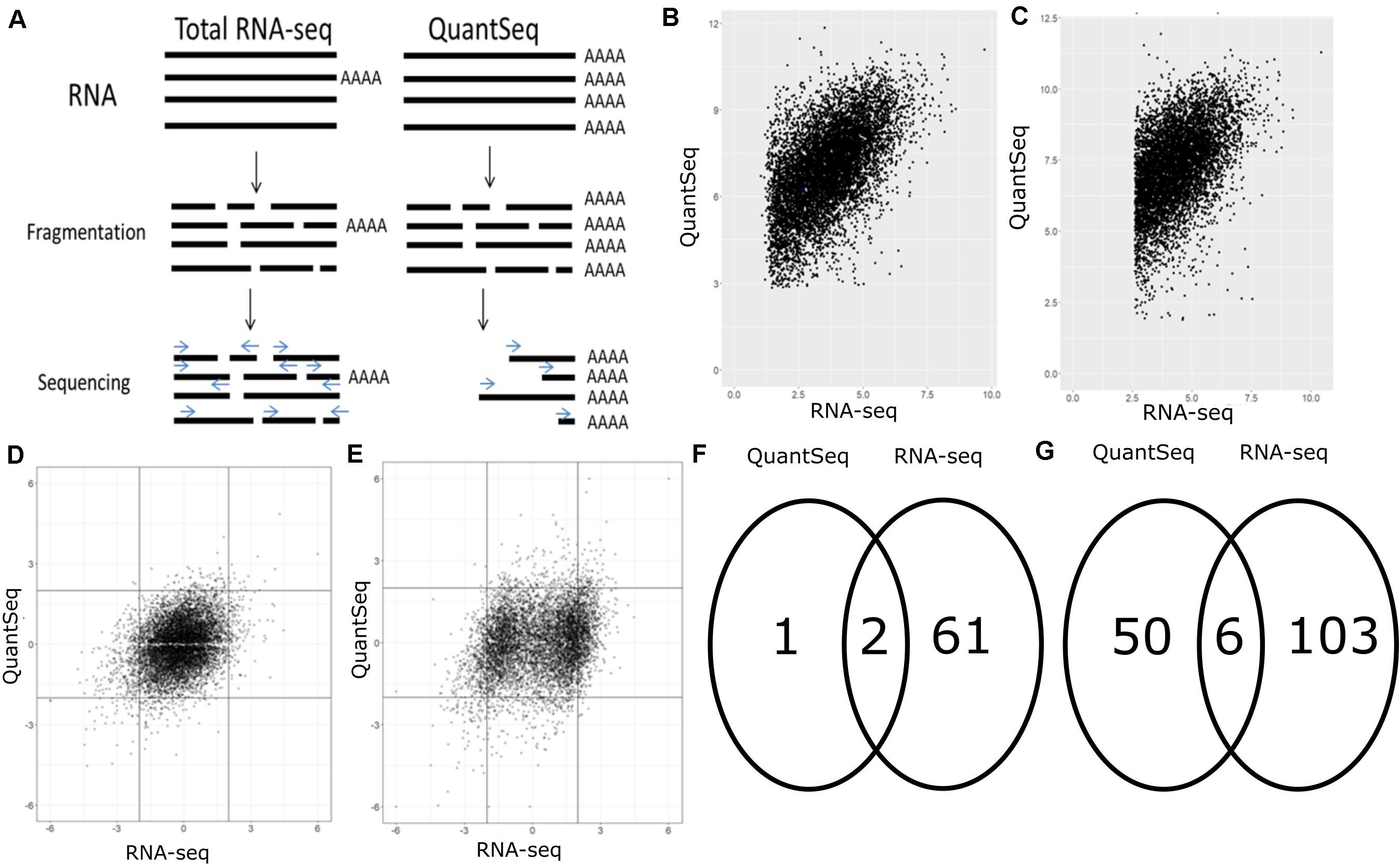
Frontiers | A Comparison of Low Read Depth QuantSeq 3′ Sequencing to Total RNA-Seq in FUS Mutant Mice