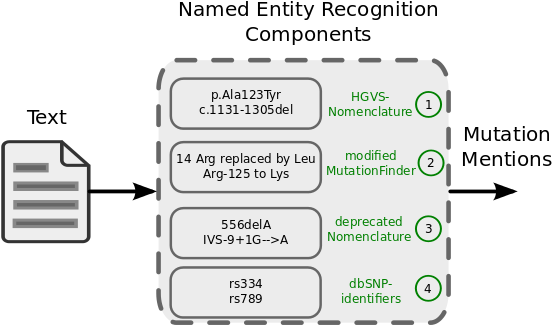
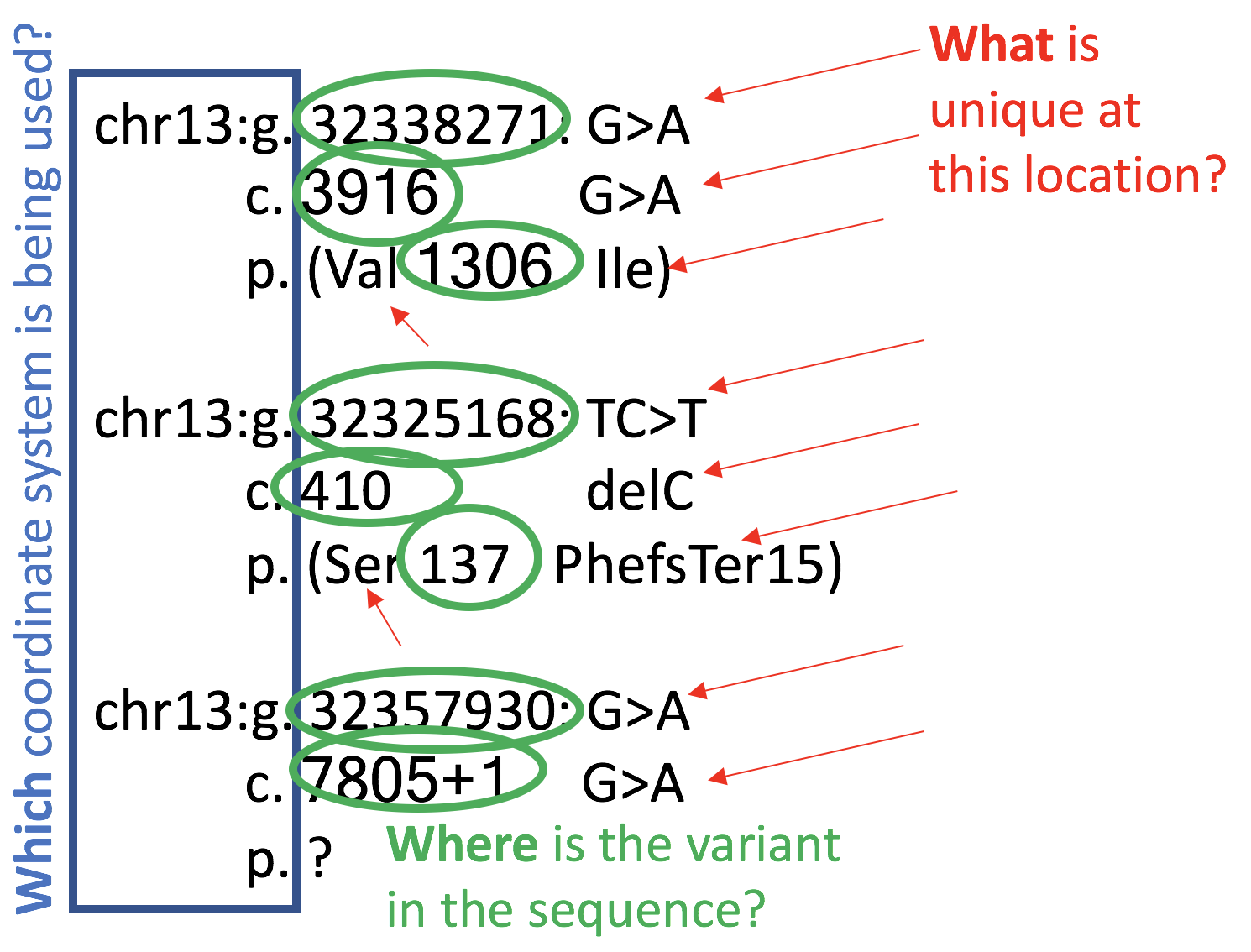
Standard mutation nomenclature in molecular diagnostics: practical and educational challenges. - Abstract - Europe PMC

Standard Mutation Nomenclature in Molecular Diagnostics: Practical and Educational Challenges - ScienceDirect

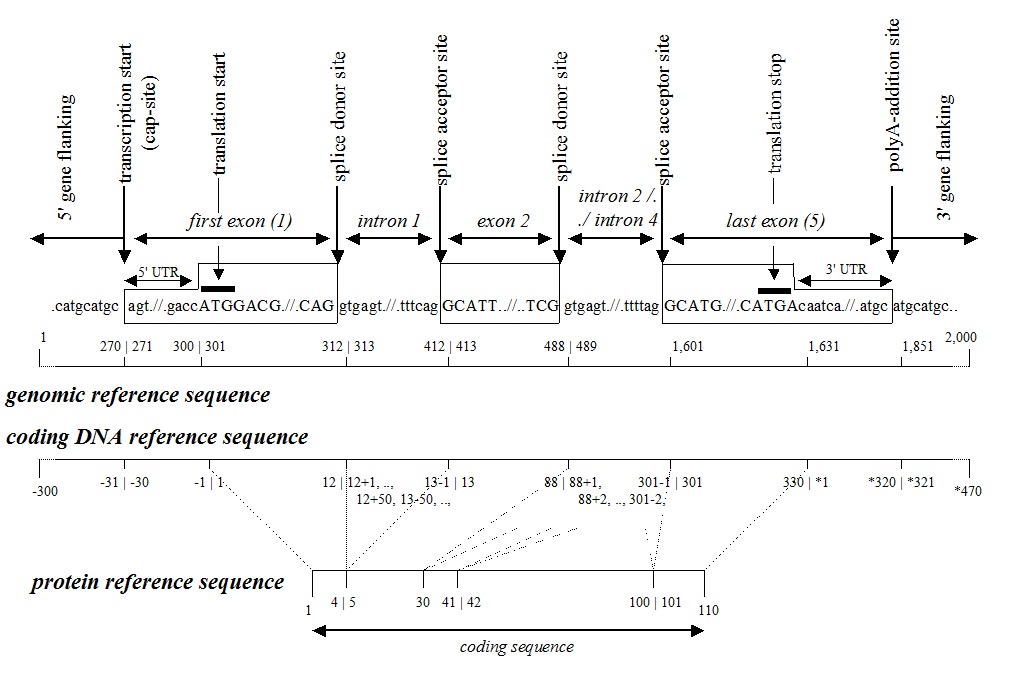
HGVS Recommendations for the Description of Sequence Variants: 2016 Update - Dunnen - 2016 - Human Mutation - Wiley Online Library
![PDF] A Python package for parsing, validating, mapping and formatting sequence variants using HGVS nomenclature | Semantic Scholar PDF] A Python package for parsing, validating, mapping and formatting sequence variants using HGVS nomenclature | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1c239b2e0b16e540ca5df6cec656132cdd821faf/2-Figure1-1.png)
PDF] A Python package for parsing, validating, mapping and formatting sequence variants using HGVS nomenclature | Semantic Scholar

A variant by any name: quantifying annotation discordance across tools and clinical databases | bioRxiv

miRNA Nomenclature: A View Incorporating Genetic Origins, Biosynthetic Pathways, and Sequence Variants: Trends in Genetics