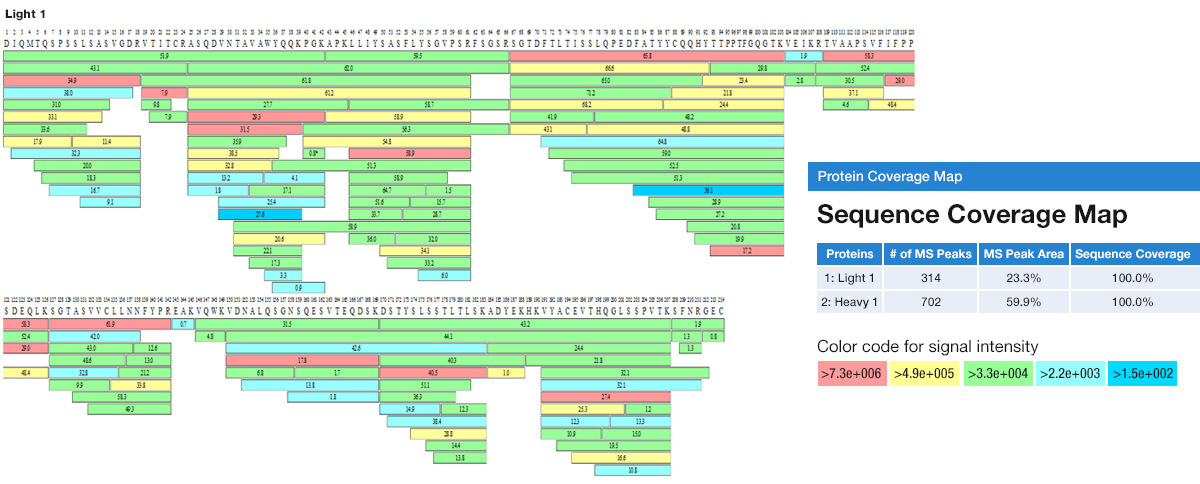
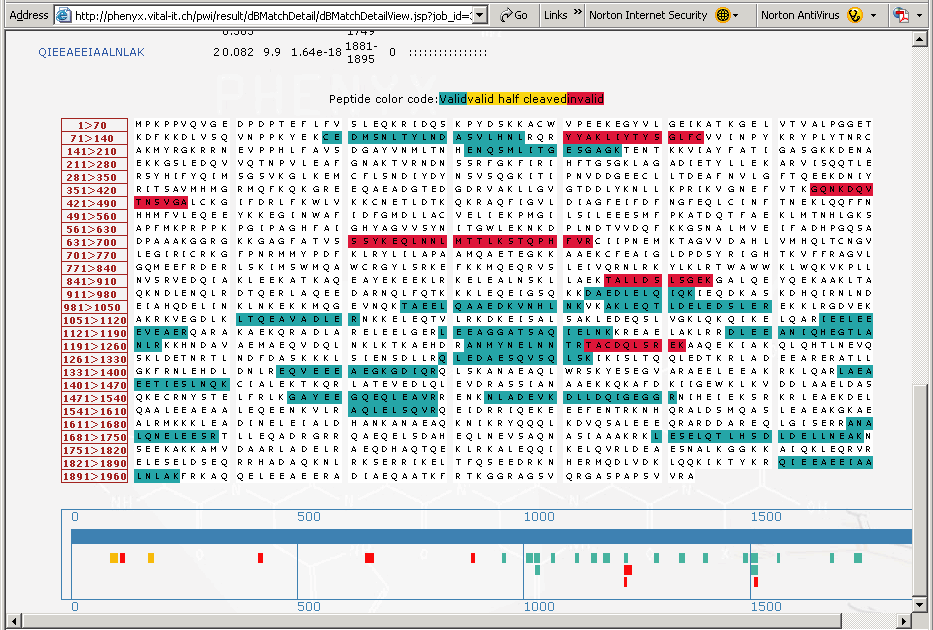
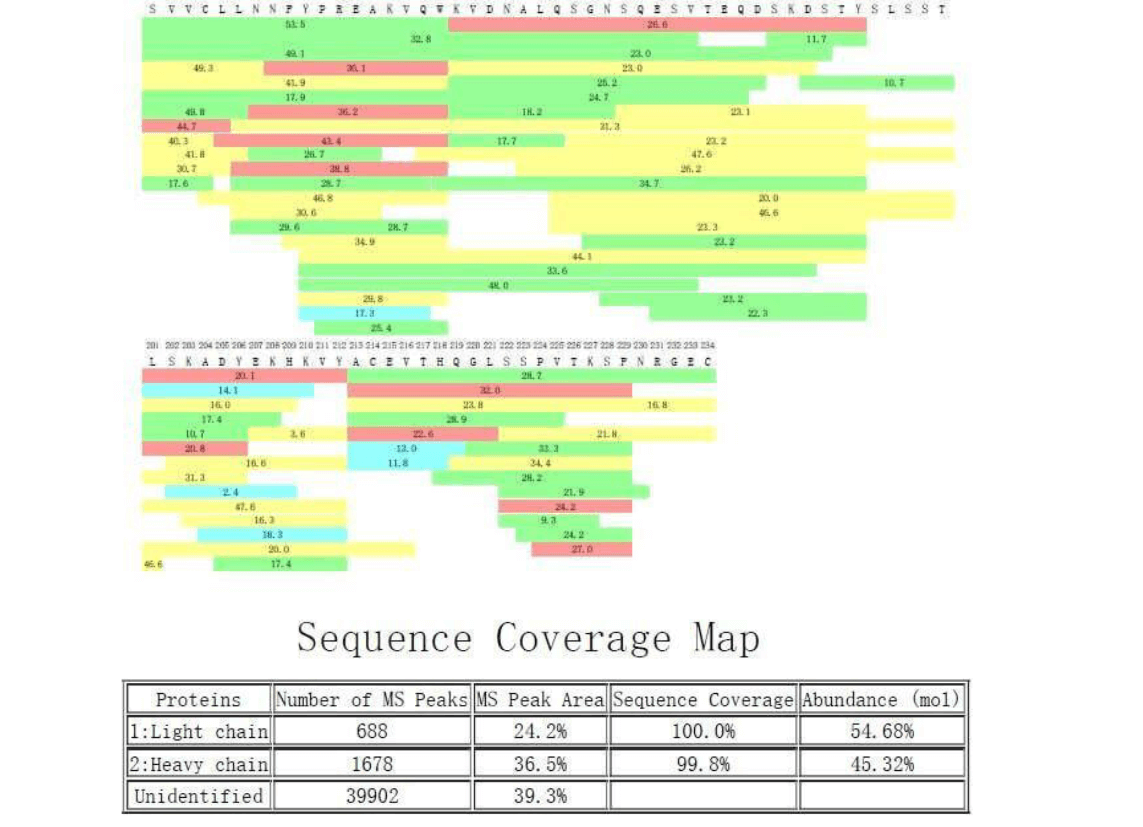
A method to identify and quantify the complete peptide composition in protein hydrolysates - ScienceDirect
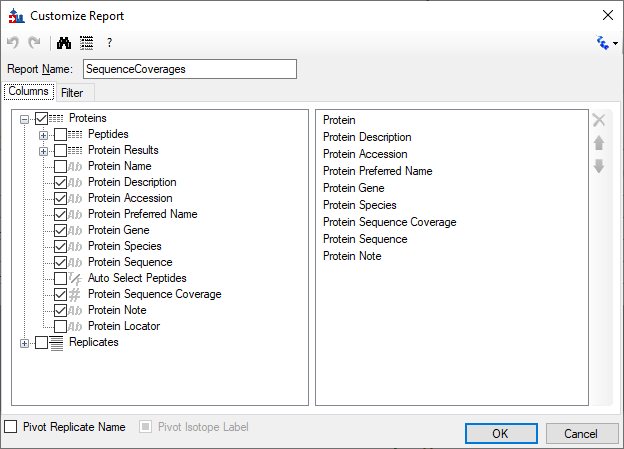
GitHub - PNNL-Comp-Mass-Spec/protein-coverage-summarizer: Computes the percent of the residues in each protein sequence that have been identified, based on a list of identified peptides. A graphical user interface (GUI) is provided to
Artificial fingerprints for cross-comparison of forensic DNA and protein recovery methods | PLOS ONE

Sequence Coverage Visualizer: A Web Application for Protein Sequence Coverage 3D Visualization | Journal of Proteome Research

Sequence Coverage Visualizer: A Web Application for Protein Sequence Coverage 3D Visualization | Journal of Proteome Research

Data-independent acquisition protease-multiplexing enables increased proteome sequence coverage across multiple fragmentation modes | bioRxiv

Maximizing Sequence Coverage in Top-Down Proteomics By Automated Multimodal Gas-Phase Protein Fragmentation | Analytical Chemistry

Figure 4. Sequence coverage of the observed protein spot svi002 peptides mapped to the known neoVTX β-subunit sequence. PMF matched peptides were shown with underlining. Further sequencing analysis was carried out by

The distribution of protein's sequence coverage, number and length of... | Download Scientific Diagram
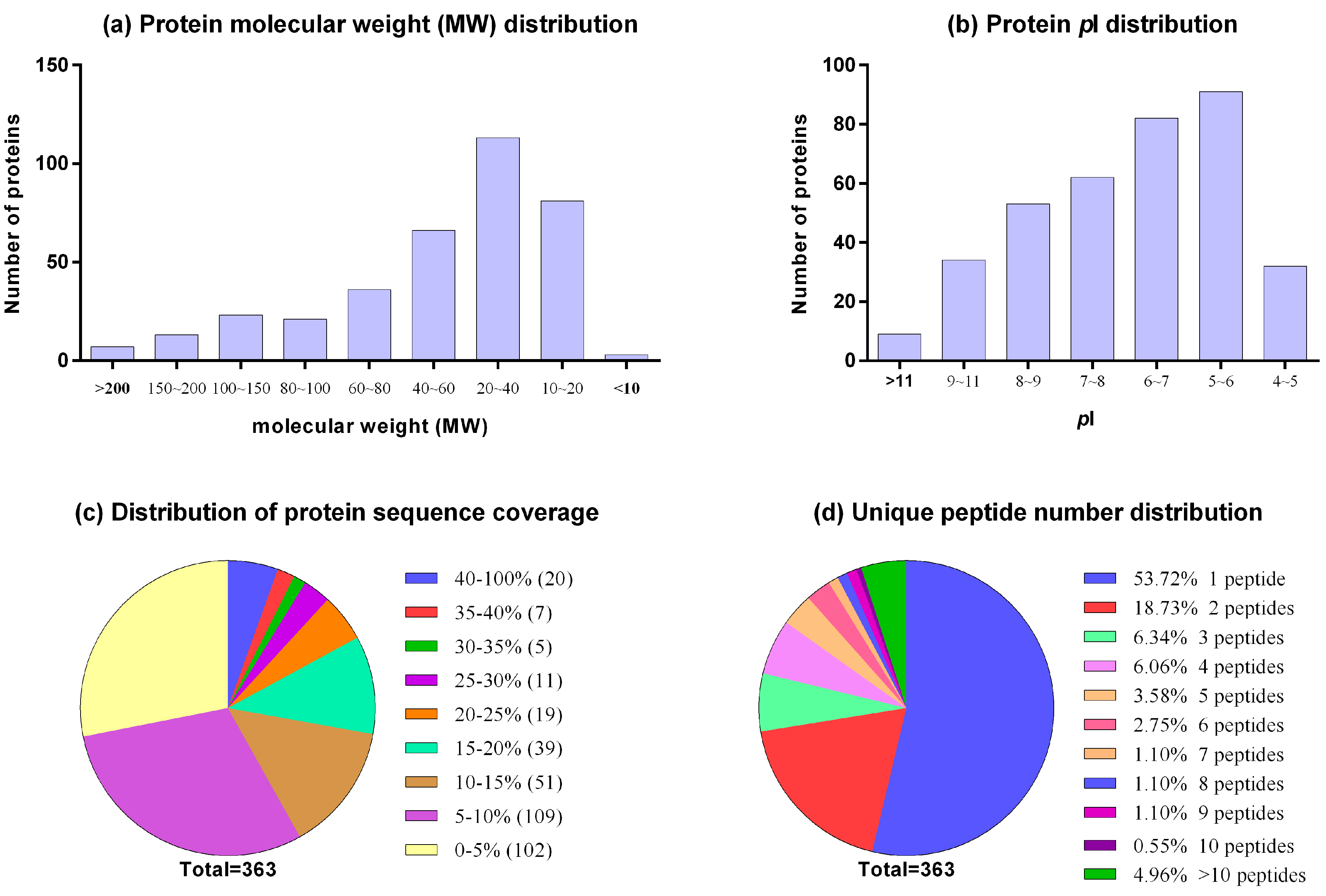
IJMS | Free Full-Text | Proteome Profile and Quantitative Proteomic Analysis of Buffalo (Bubalusbubalis) Follicular Fluid during Follicle Development

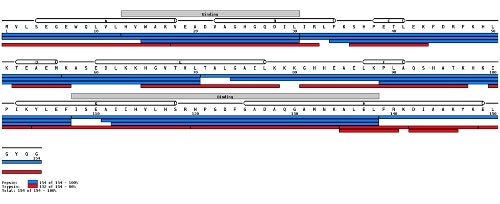
Sequence coverage view. Panel ( a ) shows heat map coverage plots for a... | Download Scientific Diagram

Internal Fragments Generated from Different Top-Down Mass Spectrometry Fragmentation Methods Extend Protein Sequence Coverage | Journal of the American Society for Mass Spectrometry

Proteome-sequence coverage for specimen Dm.5/157-16635 a, c, e, g, i,... | Download Scientific Diagram
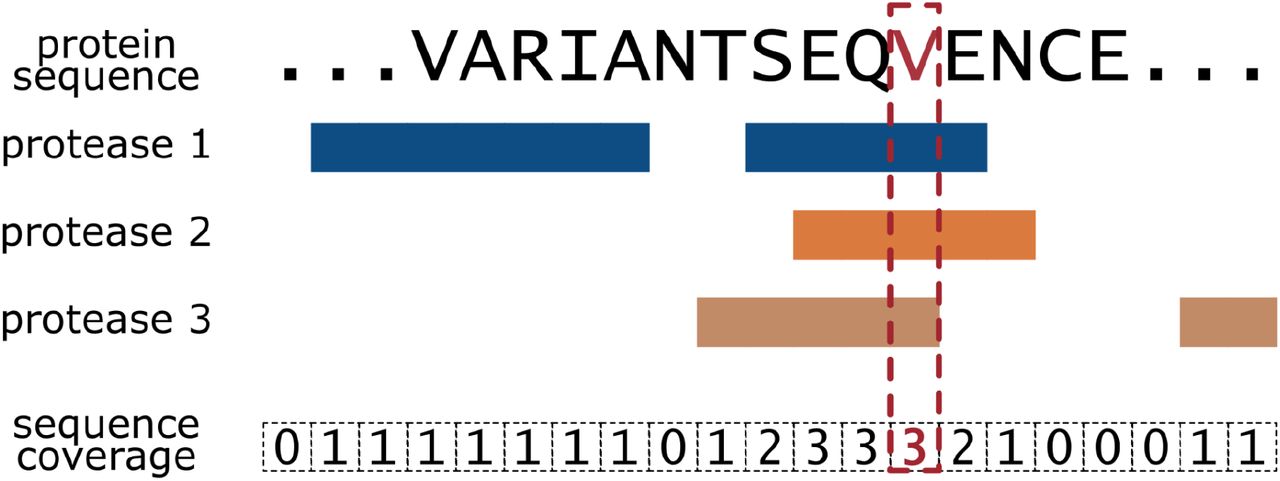
Validating amino acid variants in proteogenomics using sequence coverage by multiple reads | bioRxiv