
Claire McWhite on Twitter: "New preprint with @ProfMonaSingh. We present vcMSA, a totally new algorithm for multiple sequence alignment that's based on clustering protein language representations of amino acids. No gaps penalties,
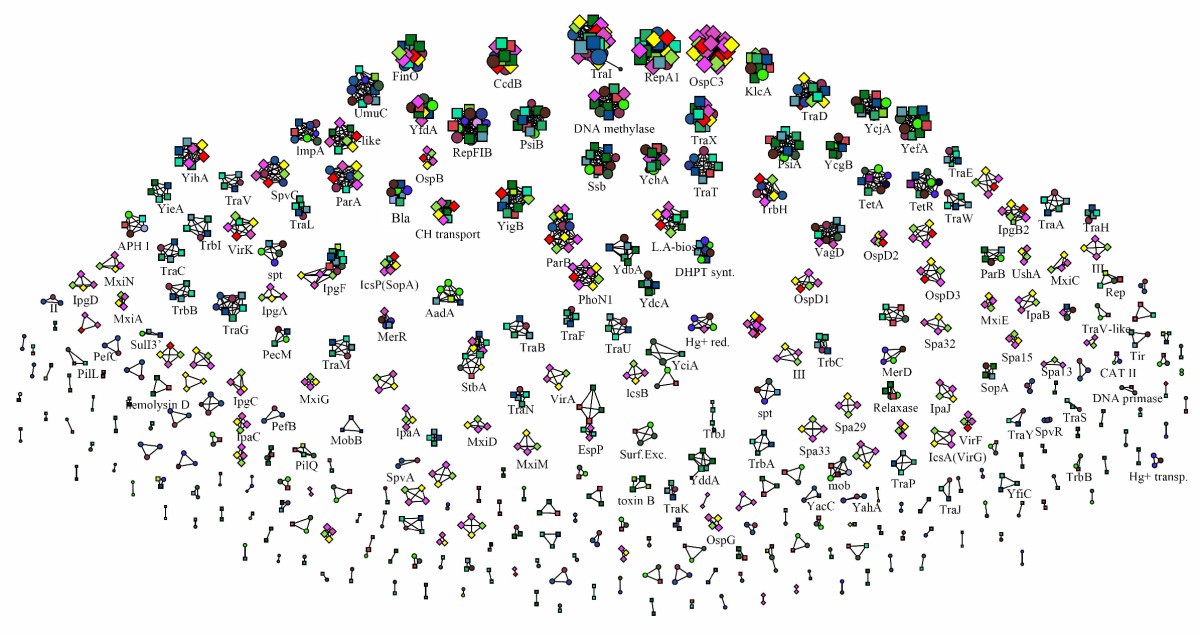

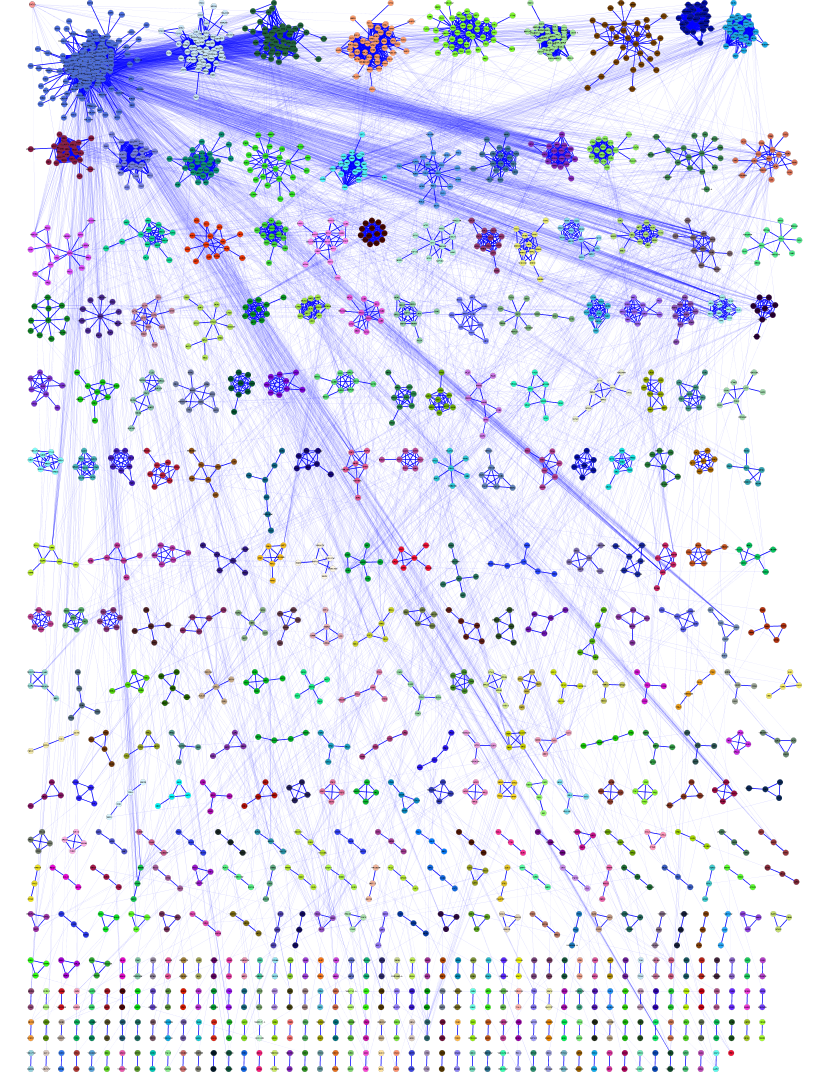
Analysis of plasmid genes by phylogenetic profiling and visualization of homology relationships using Blast2Network | BMC Bioinformatics | Full Text

Sequence similarity clustering of all MCE proteins. A representative... | Download Scientific Diagram
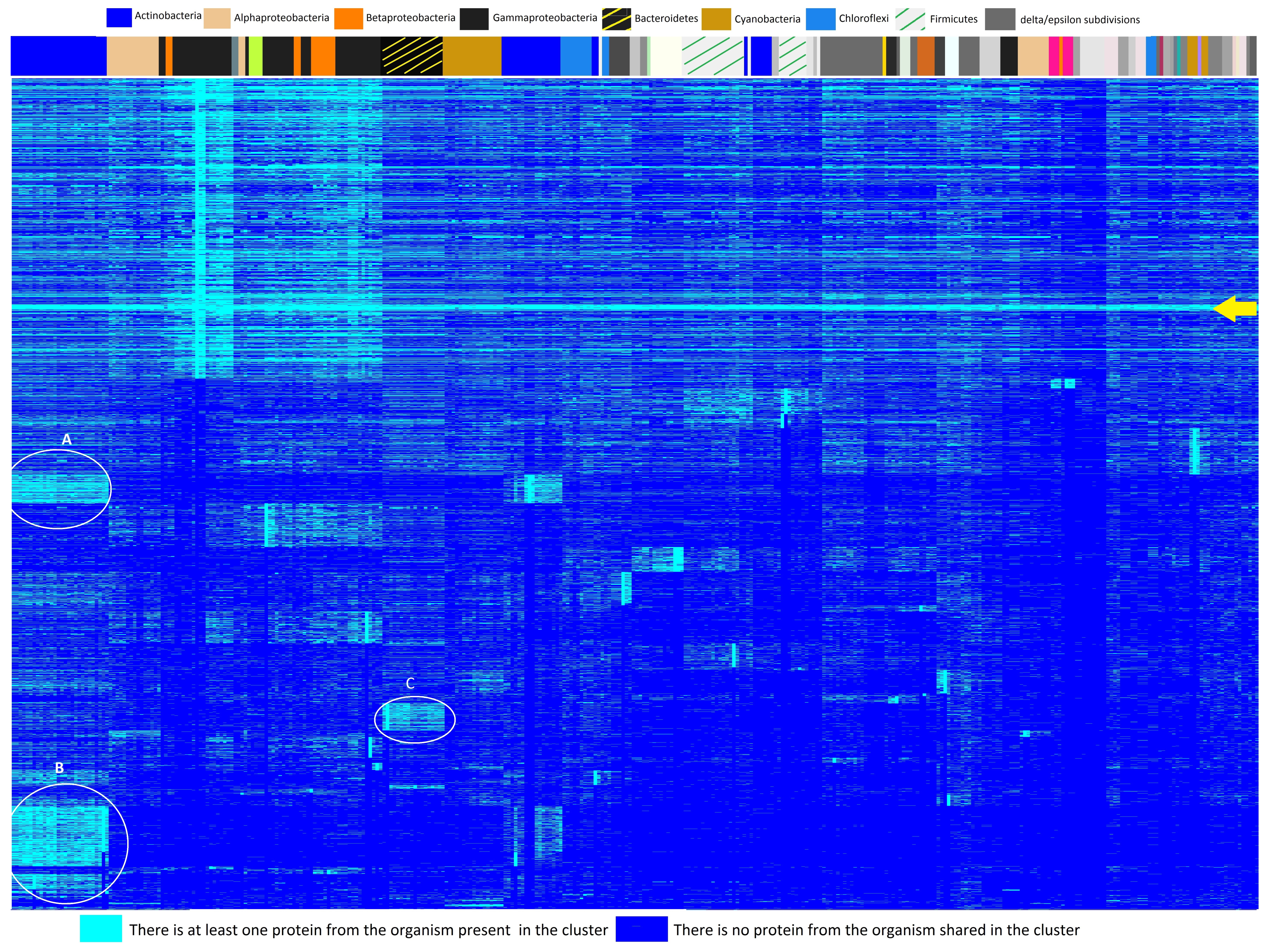
Microorganisms | Free Full-Text | A Systematic Approach to Bacterial Phylogeny Using Order Level Sampling and Identification of HGT Using Network Science

Development of a novel clustering tool for linear peptide sequences - Dhanda - 2018 - Immunology - Wiley Online Library

Protein sequence clustering has been widely used as a part of the analysis of protein structure and function. We demonstrate an approach to protein clustering, - ppt download

Large-scale sequence similarity analysis reveals the scope of sequence and function divergence in PilZ domain proteins | bioRxiv
![PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e887dab367f4c77f8b70f7d7a90b8fe77747a9bf/4-Figure1-1.png)
PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar

MSA-Regularized Protein Sequence Transformer toward Predicting Genome-Wide Chemical-Protein Interactions: Application to GPCRome Deorphanization | Journal of Chemical Information and Modeling
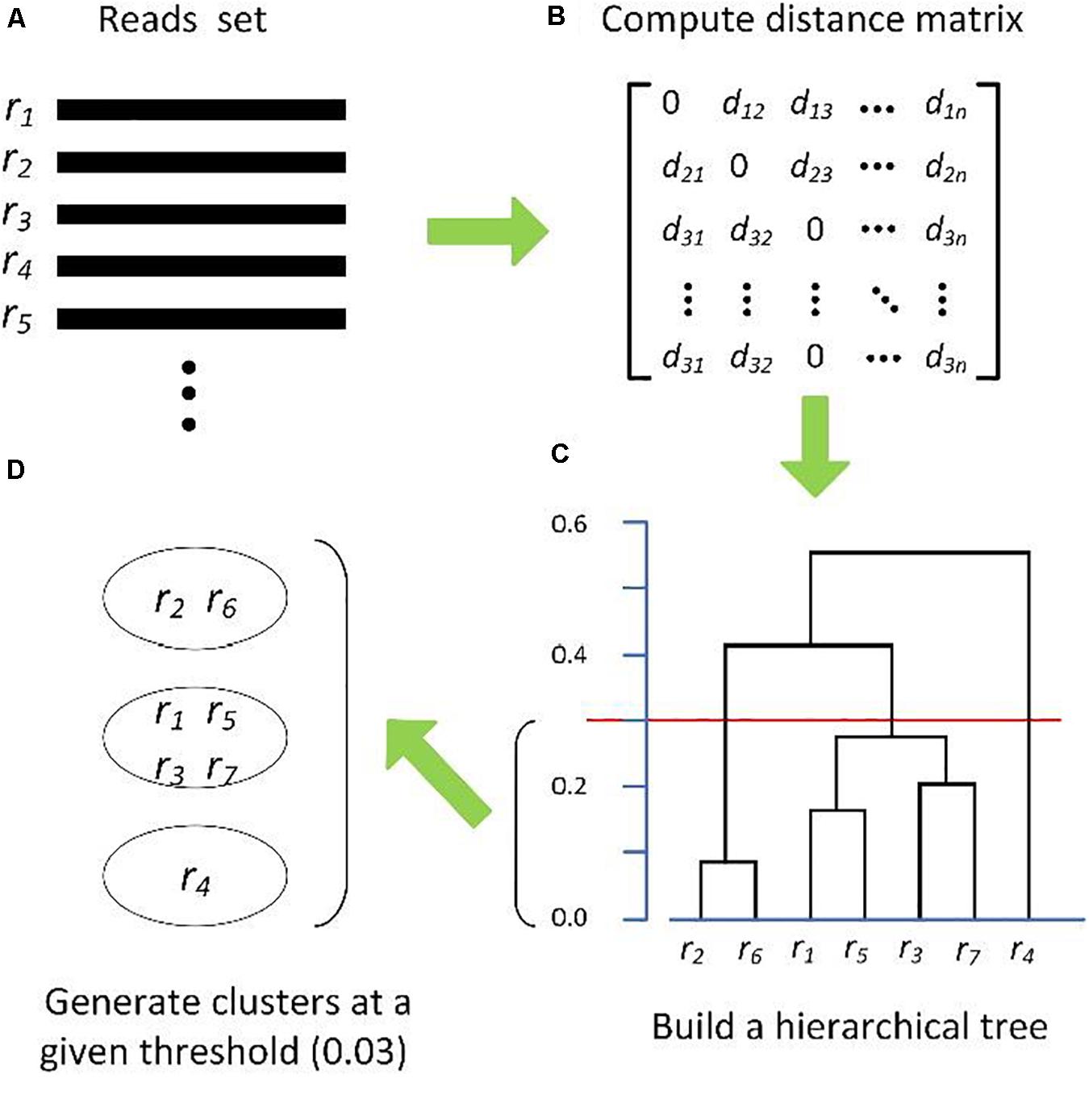
Clustering biological sequences with dynamic sequence similarity threshold | BMC Bioinformatics | Full Text
![PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f77ee5140d2c9408165c6cb756344fa6b0067f10/2-Table1-1.png)
PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar
GitHub - soedinglab/kClust: kClust is a fast and sensitive clustering method for the clustering of protein sequences. It is able to cluster large protein databases down to 20-30% sequence identity. kClust generates







![Help [PIR - Protein Information Resource] Help [PIR - Protein Information Resource]](https://proteininformationresource.org/pirwww/images/creationpirsf.jpg)