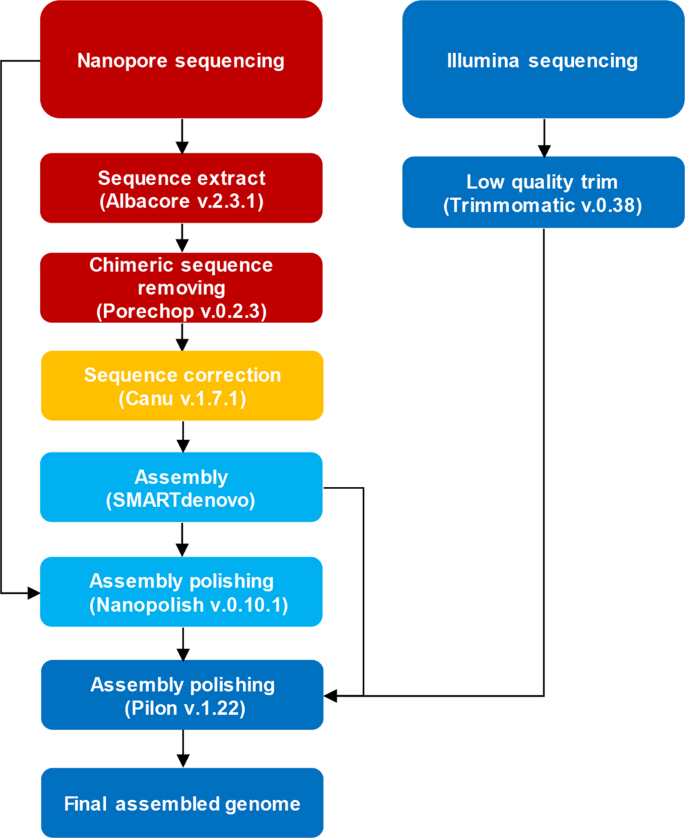
Nanopore sequencing reads improve assembly and gene annotation of the Parochlus steinenii genome | Scientific Reports
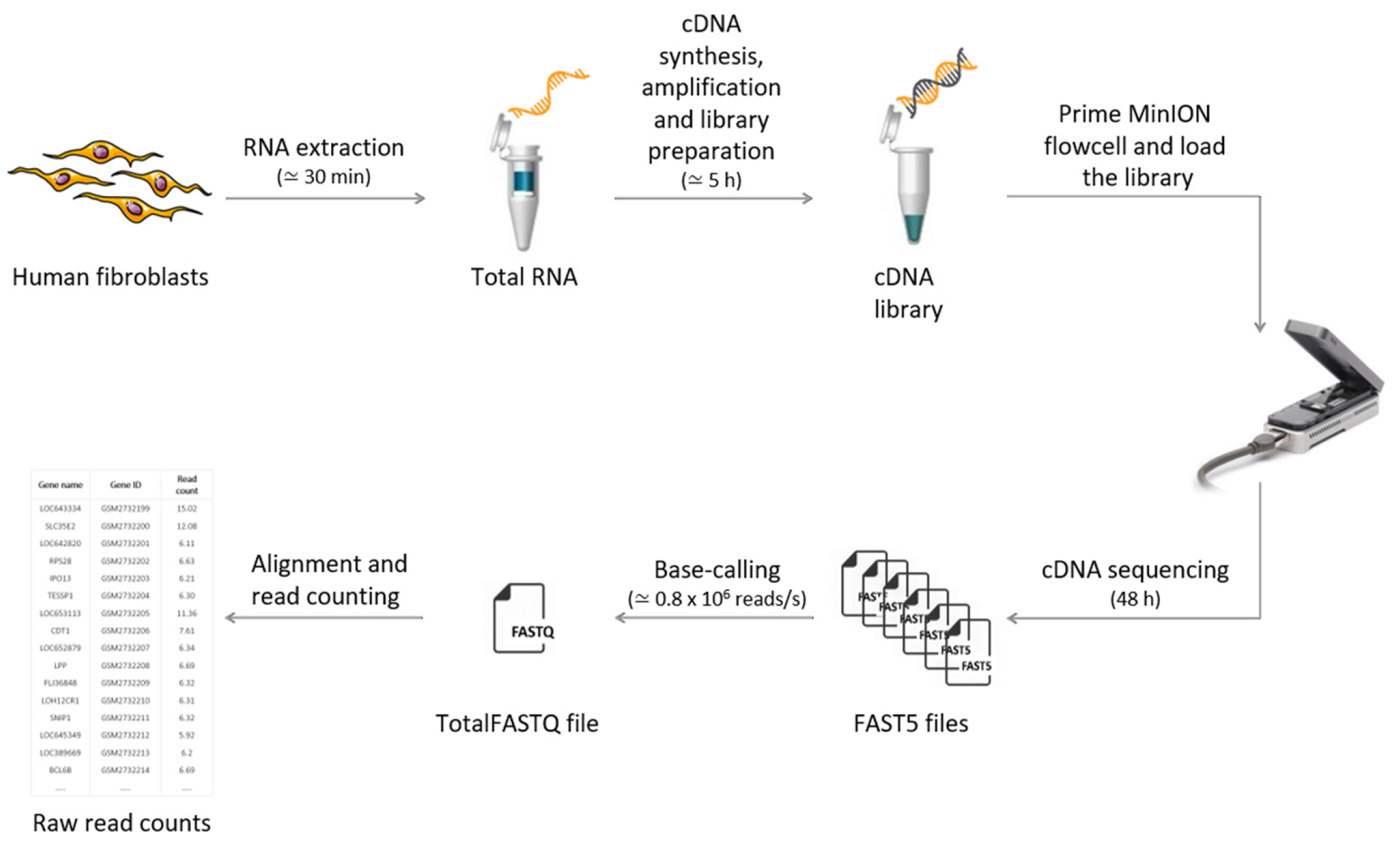
IJMS | Free Full-Text | Evaluation of Oxford Nanopore MinION RNA-Seq Performance for Human Primary Cells
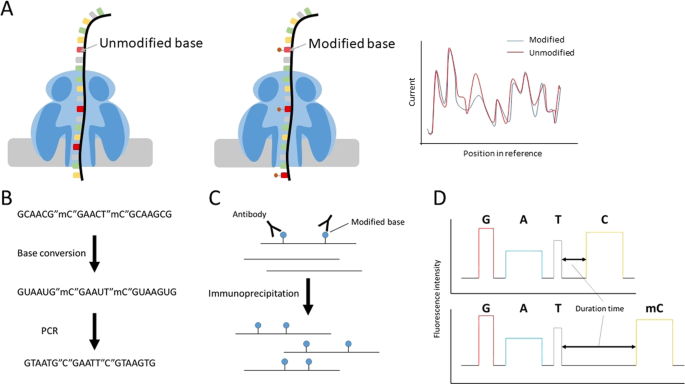
Recent advances in the detection of base modifications using the Nanopore sequencer | Journal of Human Genetics

NanoRTax, a real-time pipeline for taxonomic and diversity analysis of nanopore 16S rRNA amplicon sequencing data - ScienceDirect

Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology

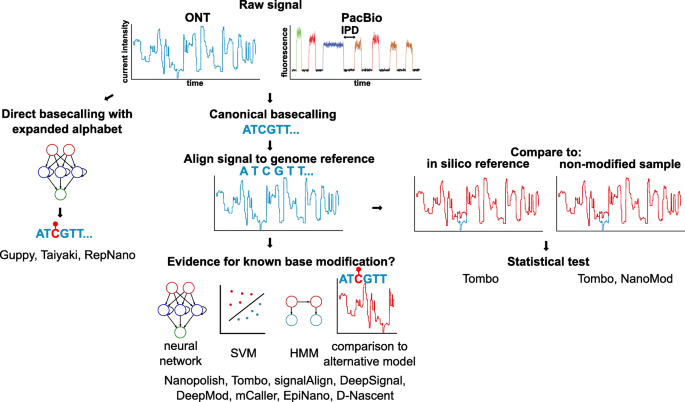
Figure 2 from Nanopore sequencing data analysis: state of the art, applications and challenges | Semantic Scholar
![PDF] Nanopore sequencing technology and tools for genome assembly: computational analysis of the current state, bottlenecks and future directions | Semantic Scholar PDF] Nanopore sequencing technology and tools for genome assembly: computational analysis of the current state, bottlenecks and future directions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1843807642bb27041970a74e6ab2cbf979f2c3bf/2-Figure1-1.png)
PDF] Nanopore sequencing technology and tools for genome assembly: computational analysis of the current state, bottlenecks and future directions | Semantic Scholar

MinION Nanopore Sequencing Data and SNV and Indel Variant Calls Obtained Using BEI Resources' Metrology Standard RNA for Zaire Mayinga Ebola Virus | IEEE DataPort
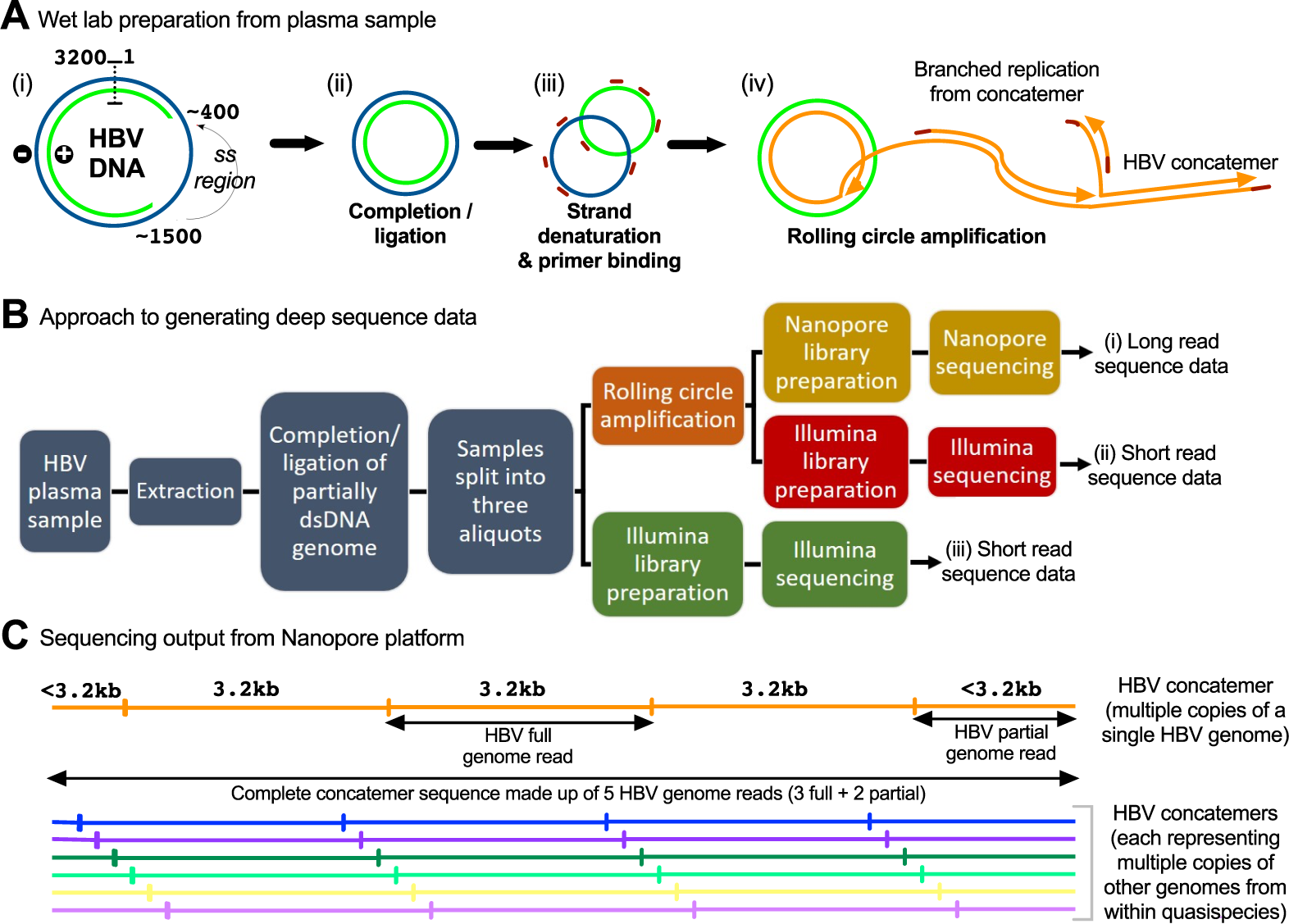
Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV) | Scientific Reports
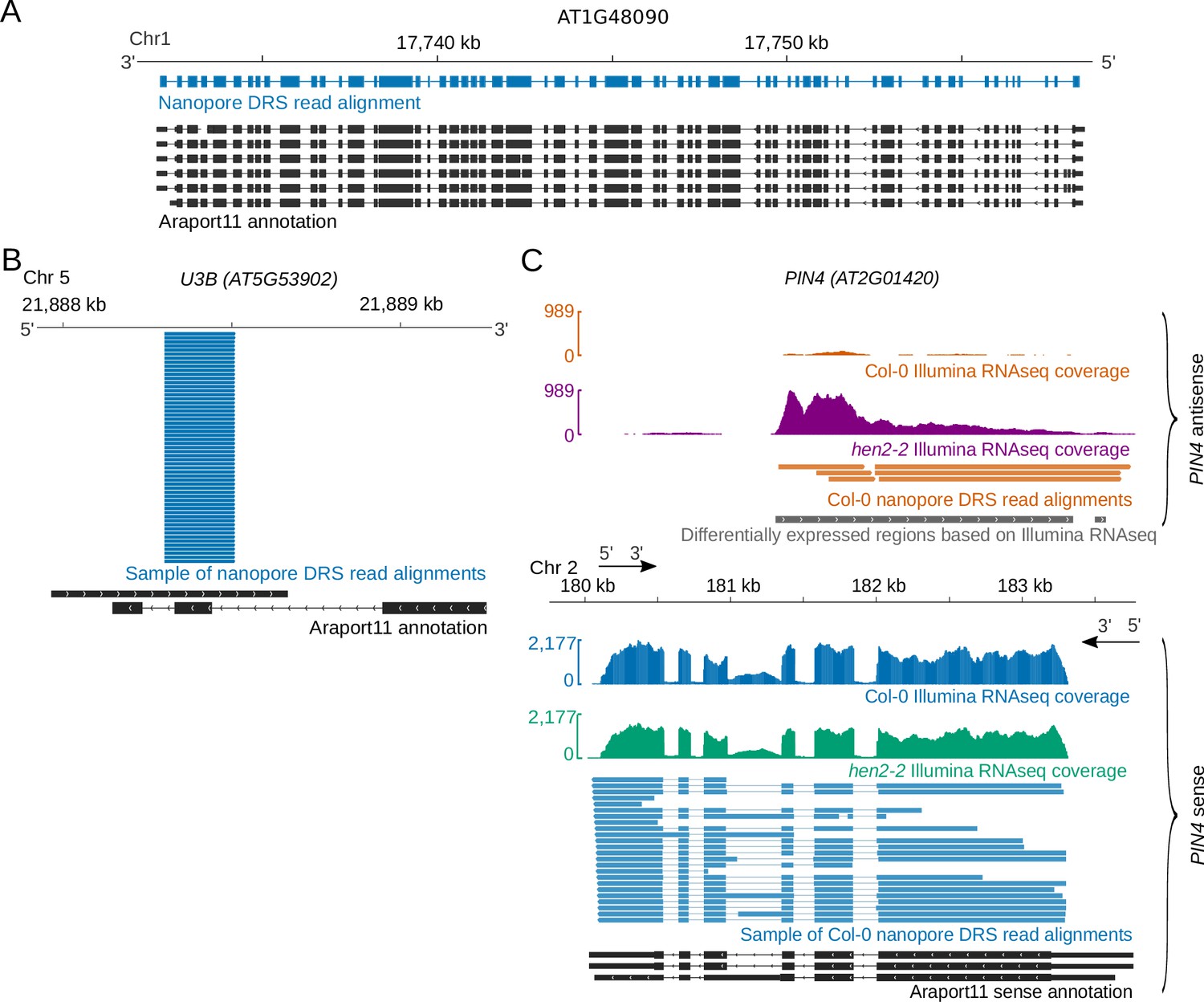
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife

Developmental validation of Oxford Nanopore Technology MinION sequence data and the NGSpeciesID bioinformatic pipeline for forensic genetic species identification - ScienceDirect