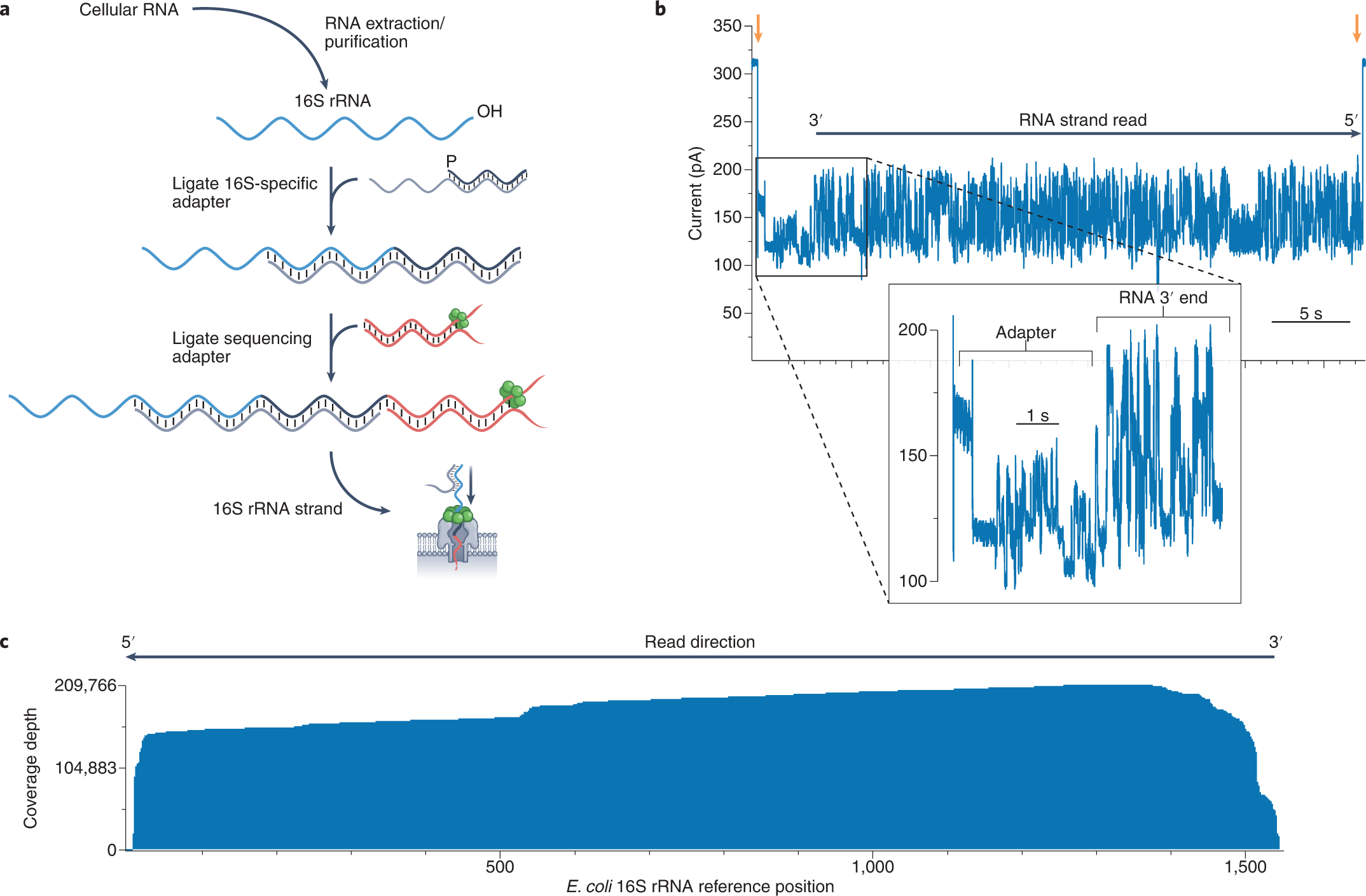
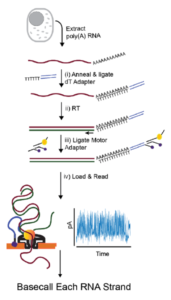
Direct detection of RNA modifications and structure using single molecule nanopore sequencing | bioRxiv
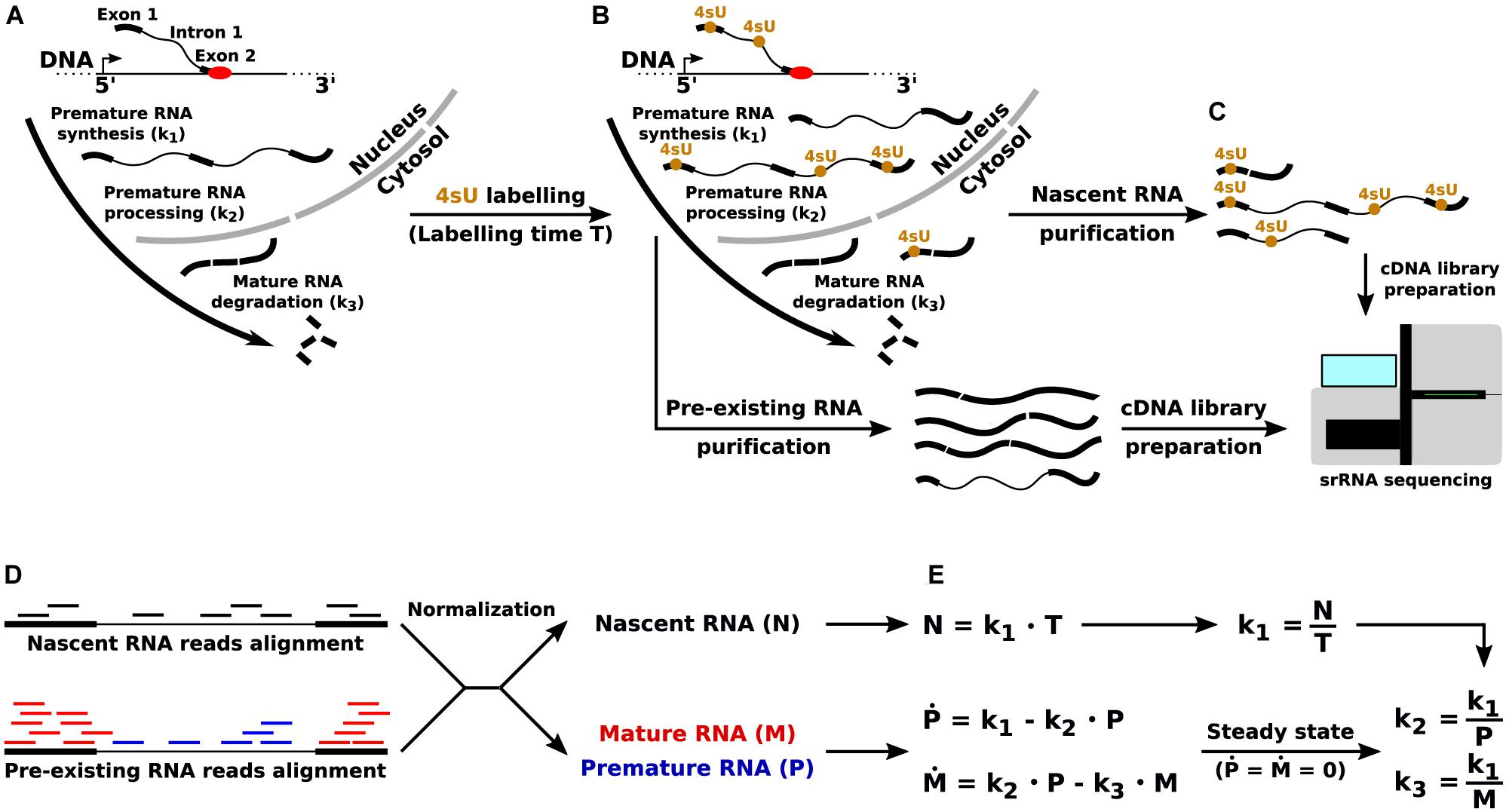
Frontiers | Direct RNA Sequencing for the Study of Synthesis, Processing, and Degradation of Modified Transcripts

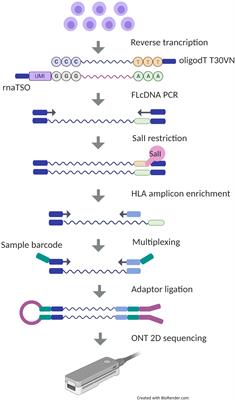
Direct RNA sequencing library preparation steps using Oxford Nanopore... | Download Scientific Diagram

Improved nanopore direct RNA sequencing of cardiac myocyte samples by selective mt-RNA depletion - Journal of Molecular and Cellular Cardiology
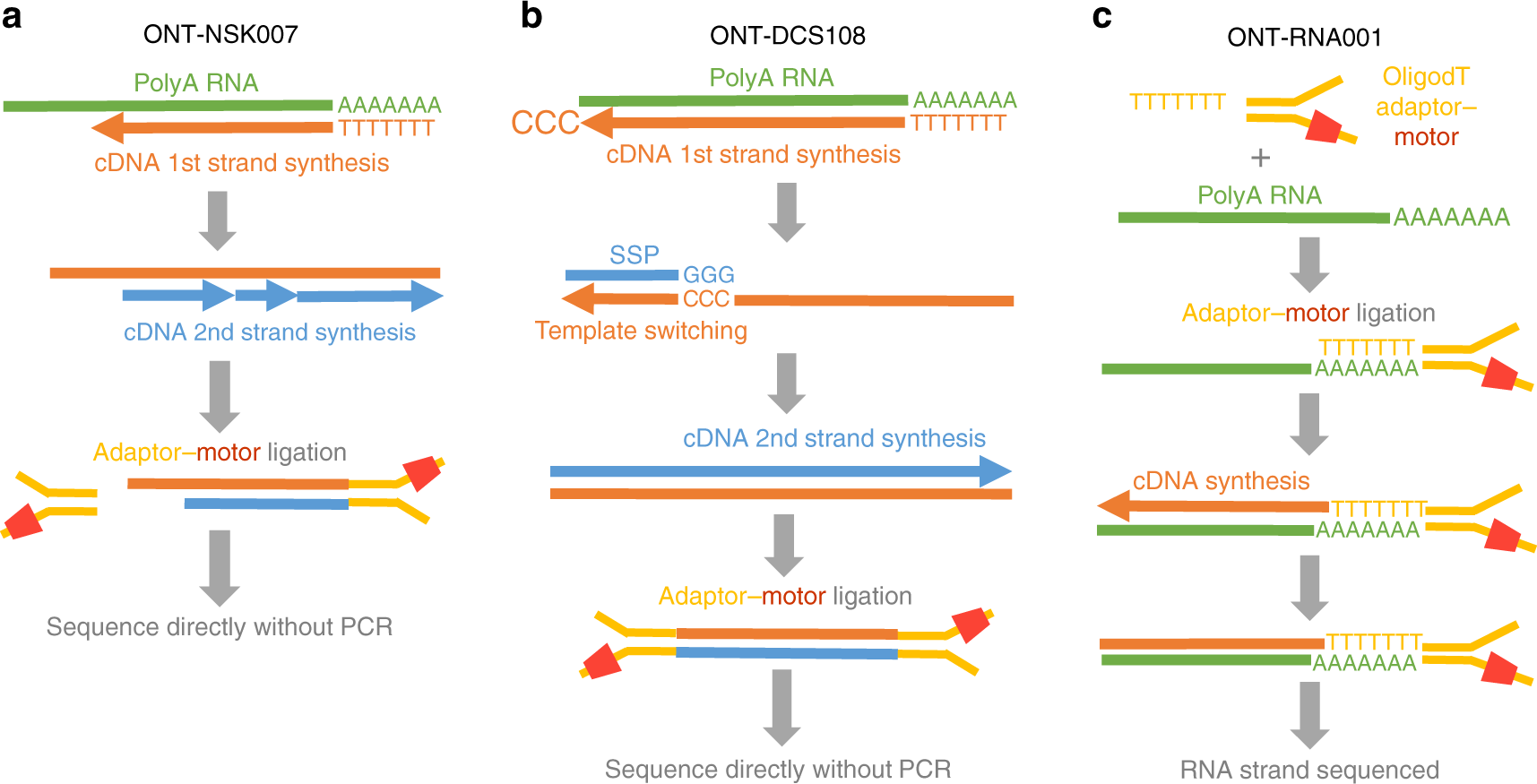
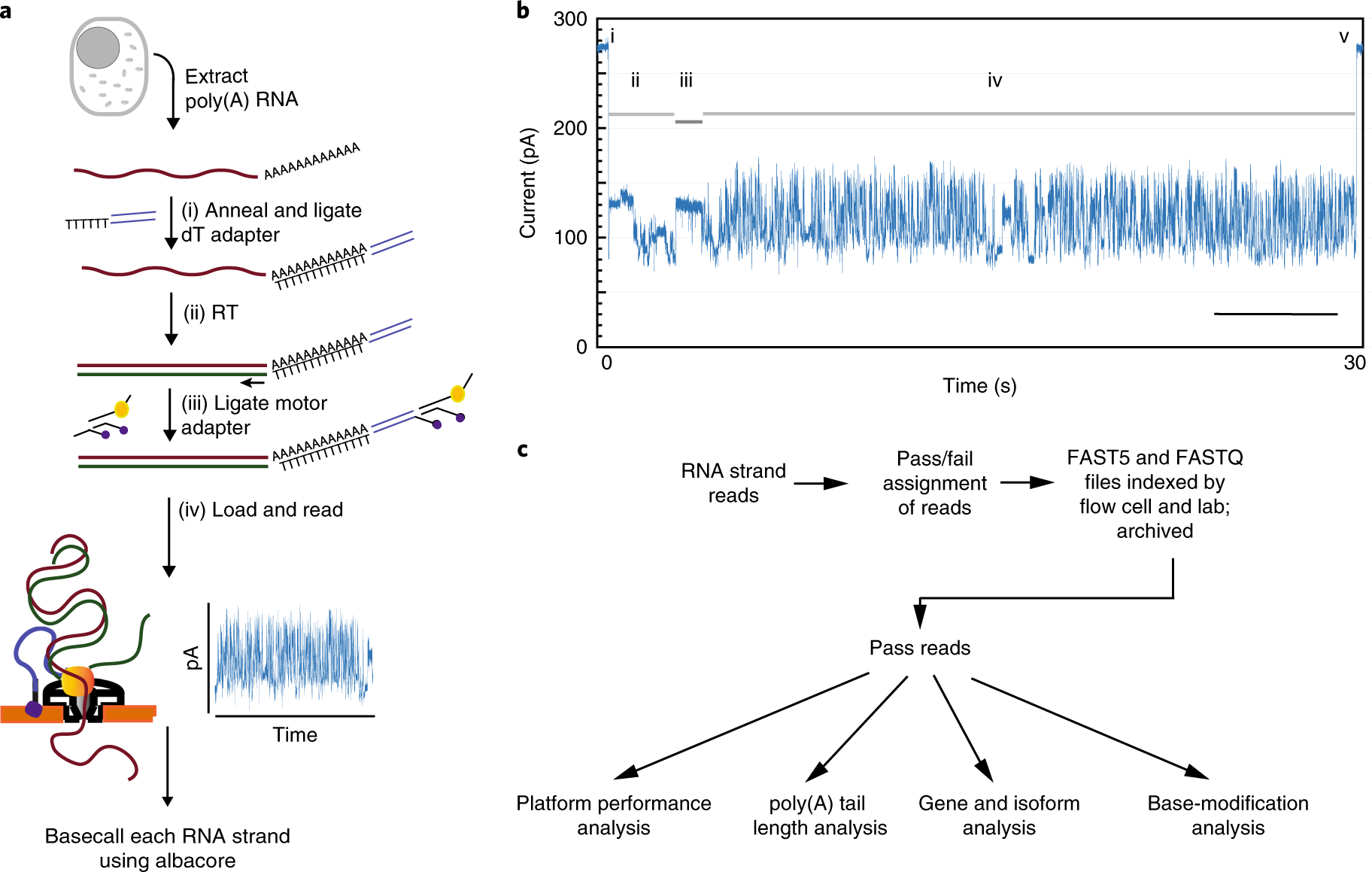
A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes | Nature Communications
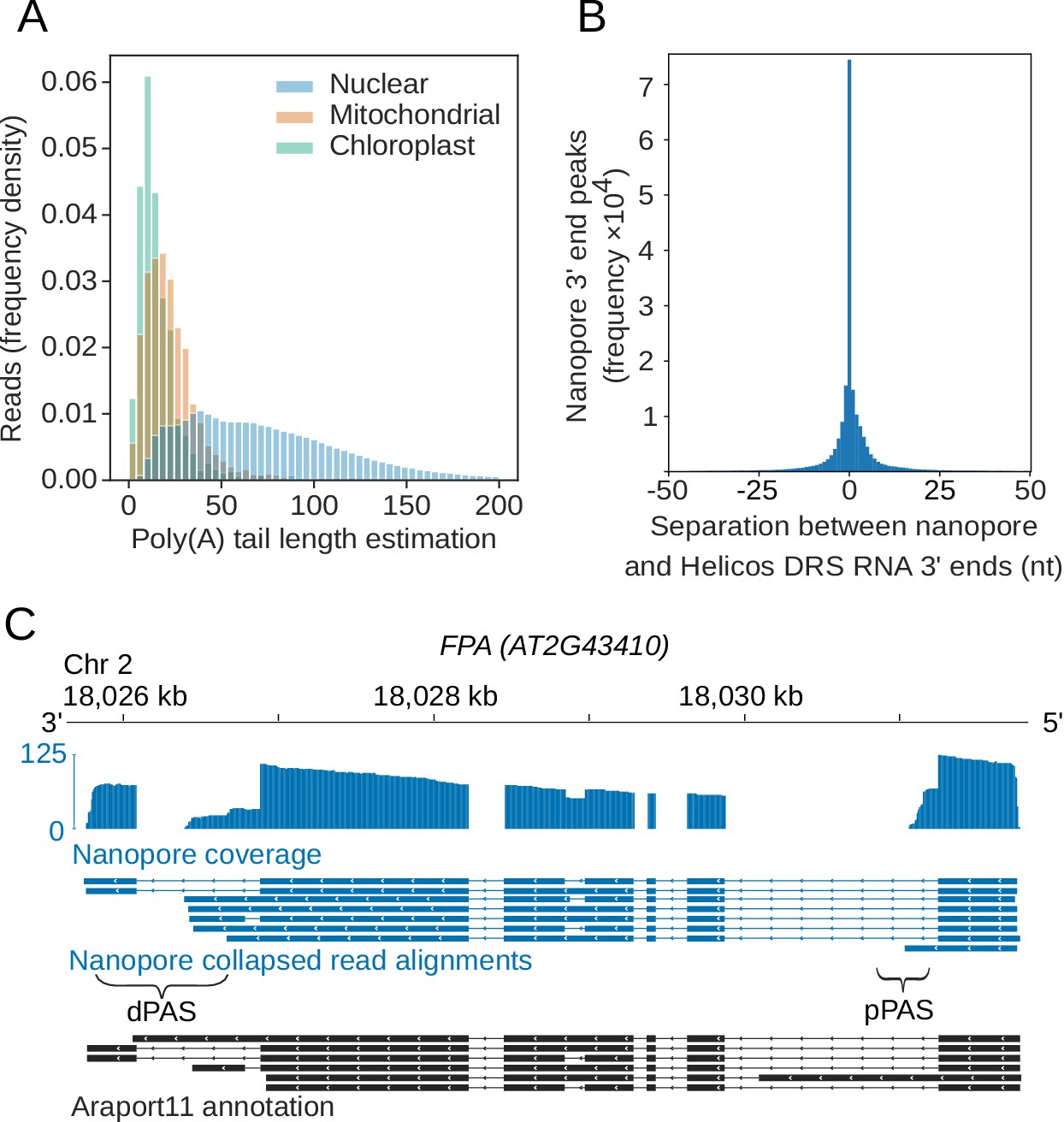
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife
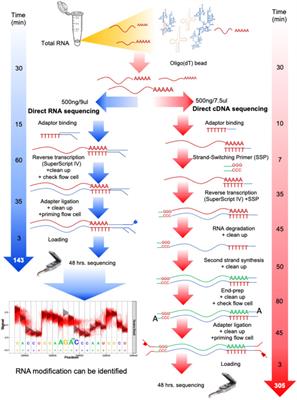
Frontiers | Native RNA or cDNA Sequencing for Transcriptomic Analysis: A Case Study on Saccharomyces cerevisiae
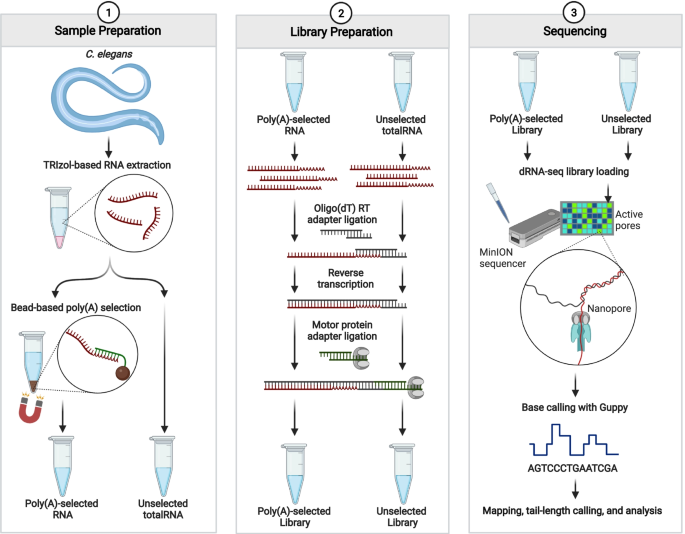
Poly(a) selection introduces bias and undue noise in direct RNA-sequencing | BMC Genomics | Full Text

Analysis of Transcriptome and Epitranscriptome in Plants Using PacBio Iso- Seq and Nanopore-Based Direct RNA Sequencing. - Abstract - Europe PMC

Barcoding and demultiplexing Oxford Nanopore native RNA sequencing reads with deep residual learning | bioRxiv

Beyond sequencing: machine learning algorithms extract biology hidden in Nanopore signal data: Trends in Genetics

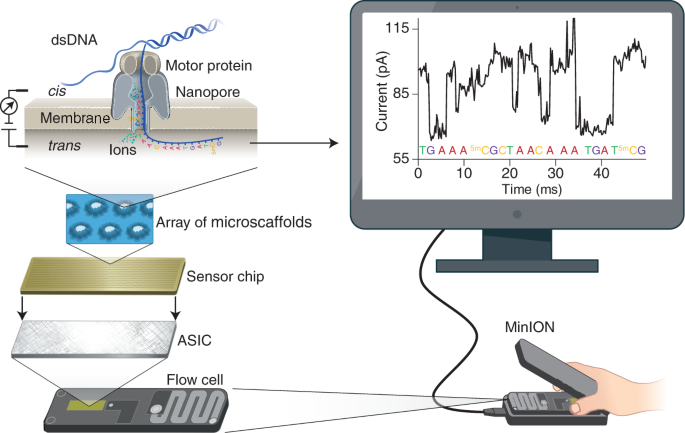
Recent advances in the detection of base modifications using the Nanopore sequencer | Journal of Human Genetics