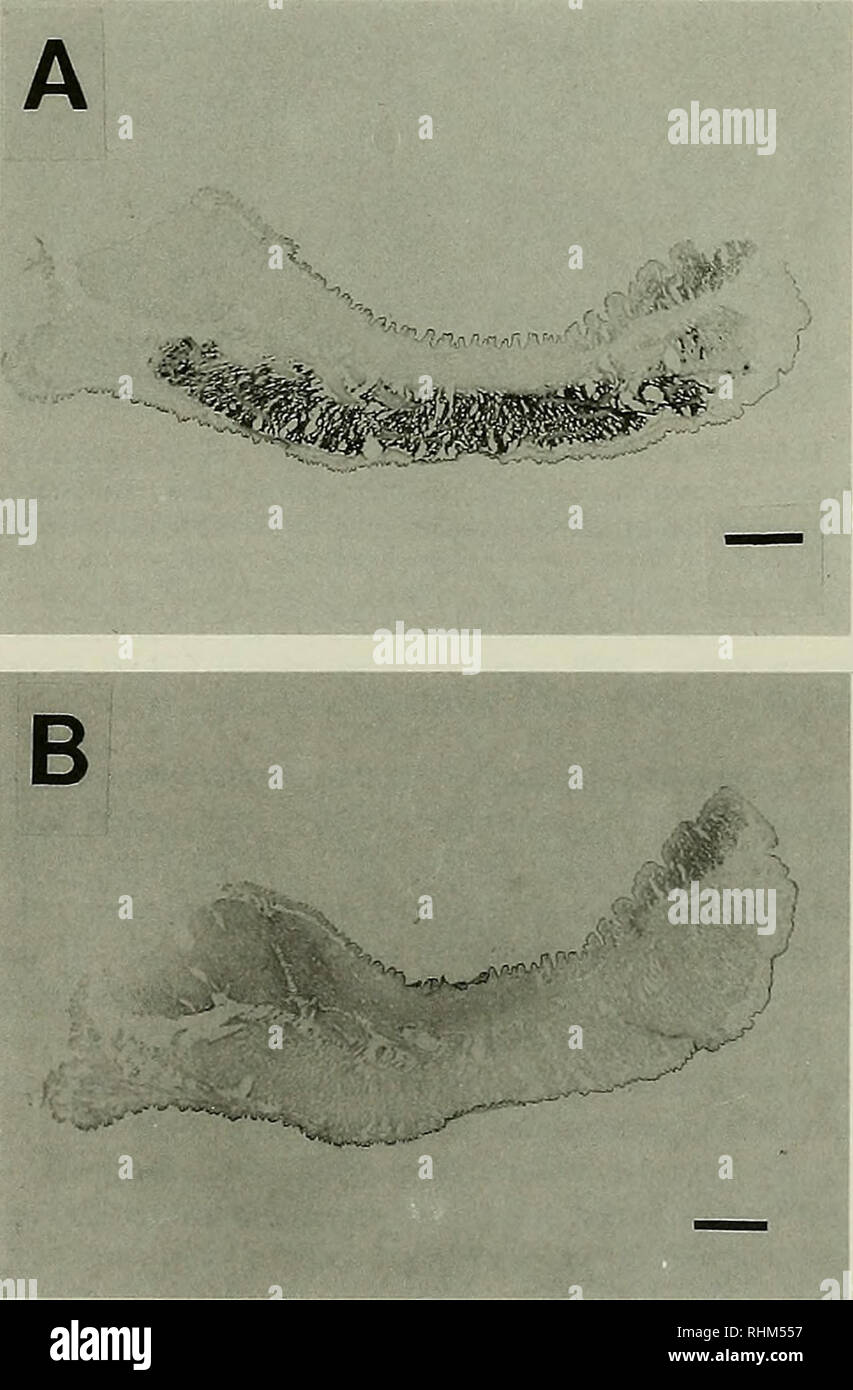
The Biological bulletin. Biology; Zoology; Marine biology. Figure 2. Northern blot hybridization on RNA extracted from the distal part and the rest of the foot of Mytihis gallopmvincialis. Ten mi- crograms

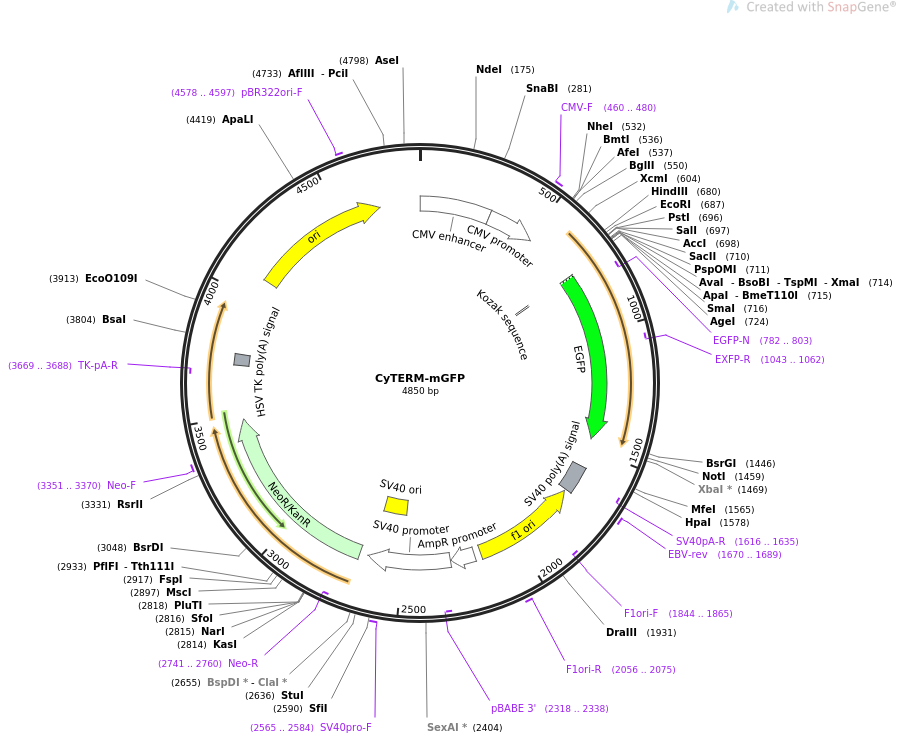
Schematic of pBIN19-mGFP-ER and Tn5393 and their integration site in... | Download Scientific Diagram
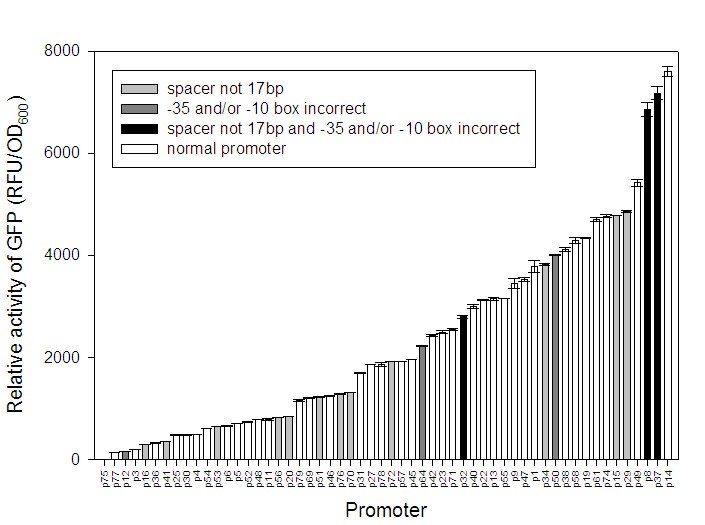
Construction and model-based analysis of a promoter library for E. coli: an indispensable tool for metabolic engineering | BMC Biotechnology | Full Text

Antiparallel dimer structure of CELSR cadherin in solution revealed by high-speed atomic force microscopy | PNAS

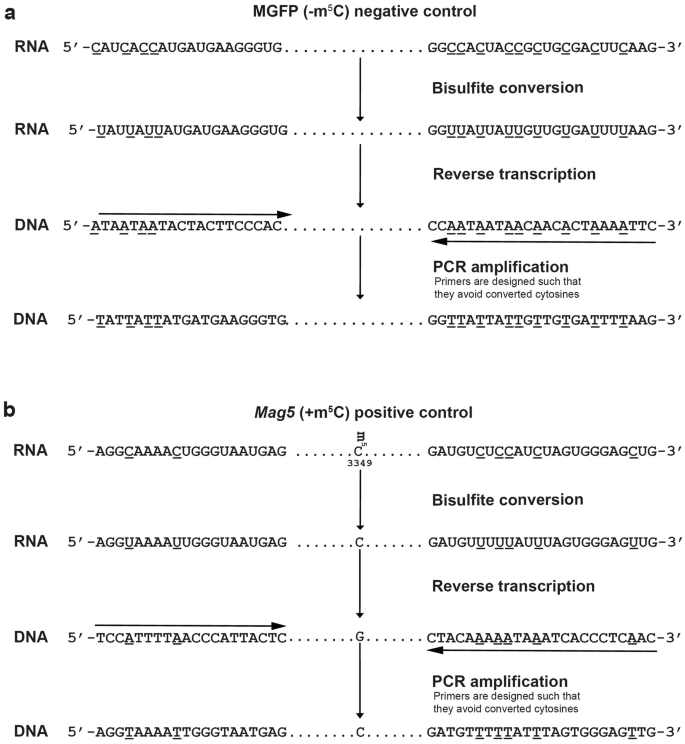
Expression of functional recombinant mussel adhesive protein Mgfp-5 in Escherichia coli. - Abstract - Europe PMC

An intramolecular scrambling path controlled by a gatekeeper in Xkr8 phospholipid scramblase | bioRxiv

Sequence alignments for nucleotides (A) and amino acids (B) of Mgfp-5... | Download Scientific Diagram