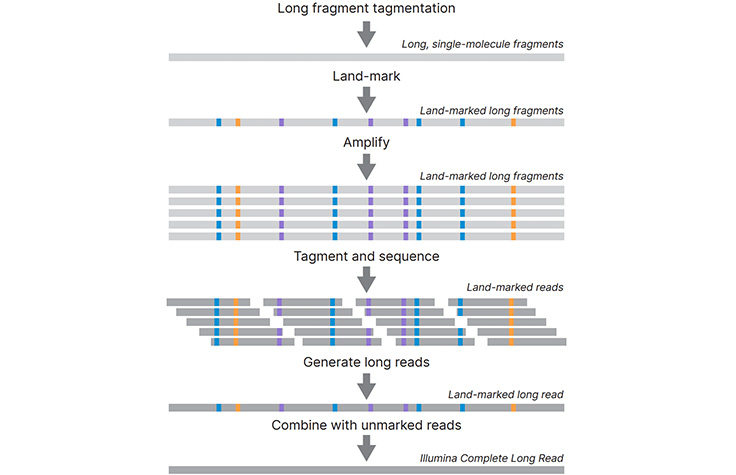
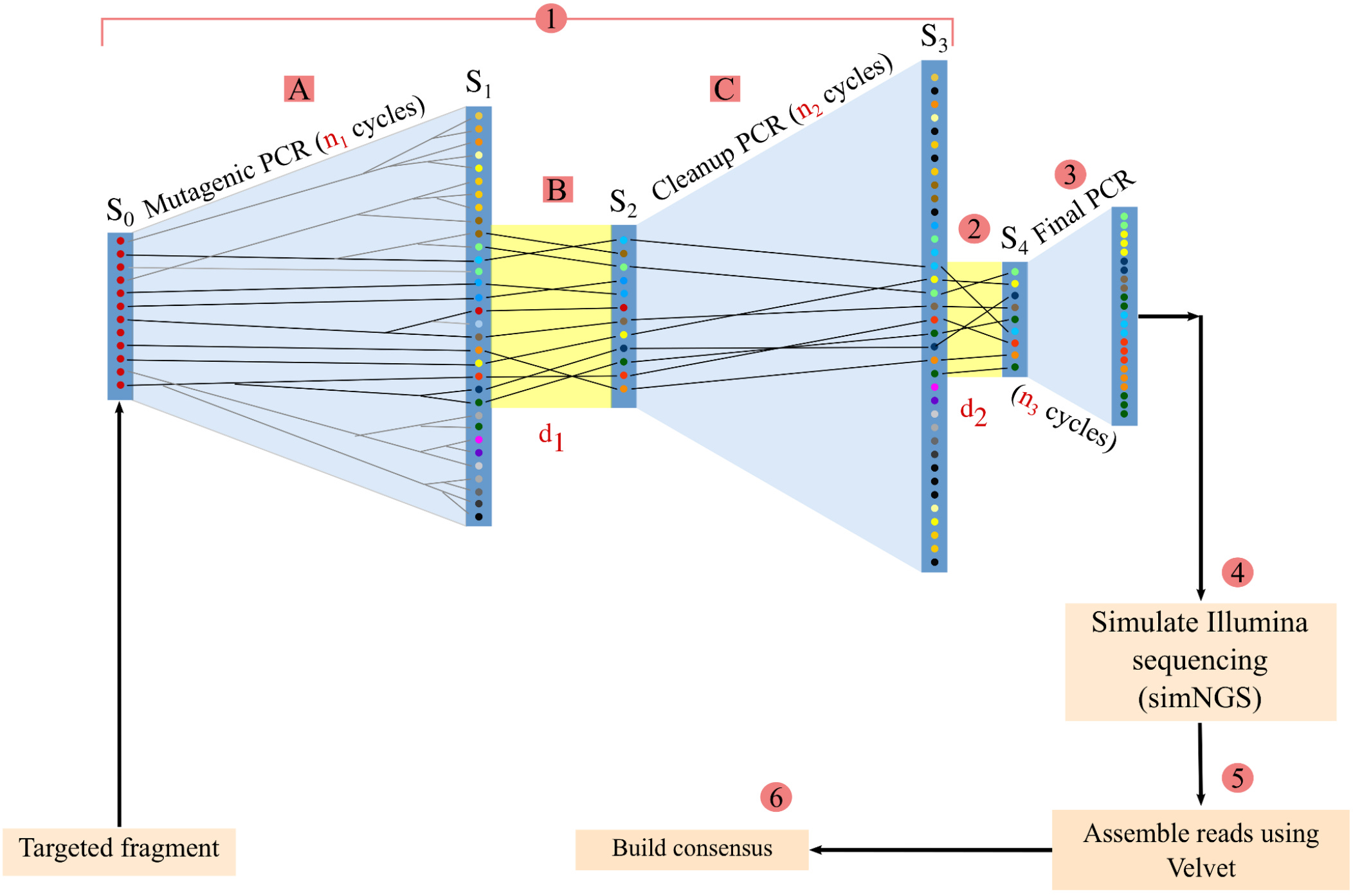
Nick Loman on Twitter: "@koadman @longastech @torstenseemann Introducing Morphoseq: a way of getting long "virtual" reads from short read platforms like Illumina. The basic principle is to mutagenise sample to actually remove

Longas introduces DNA sequencing technology for virtual long read capability + | Bioworld | BioWorld

Nick Loman on Twitter: "@koadman @longastech @torstenseemann Introducing Morphoseq: a way of getting long "virtual" reads from short read platforms like Illumina. The basic principle is to mutagenise sample to actually remove
Long Read Sequencing Market is expected to reach US$ 5,334.68 million by 2028 With Oxford Nanopore Technologies; Tataa Biocenter; Illumina, Inc; Perkinelmer Inc.; F. Hoffmann-La Roche Ltd.; Baseclear B.V.; Bionano Genomics; Longas

Nick Loman on Twitter: "@koadman @longastech @torstenseemann Introducing Morphoseq: a way of getting long "virtual" reads from short read platforms like Illumina. The basic principle is to mutagenise sample to actually remove
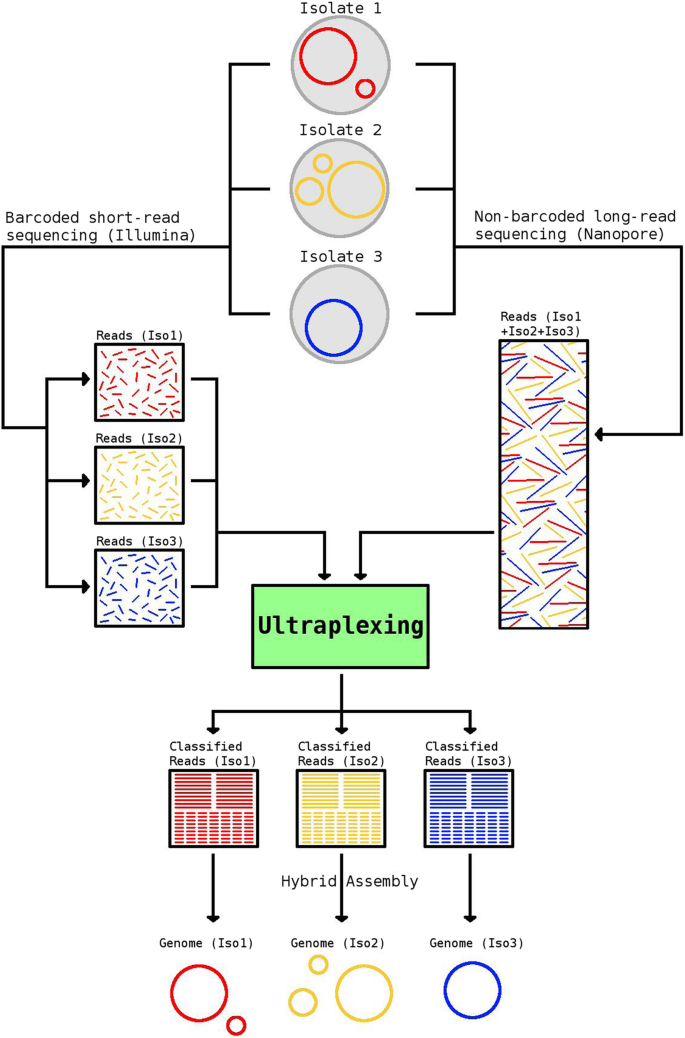
Ultraplexing: increasing the efficiency of long-read sequencing for hybrid assembly with k-mer-based multiplexing | Genome Biology | Full Text

Nick Loman on Twitter: "@koadman @longastech @torstenseemann Introducing Morphoseq: a way of getting long "virtual" reads from short read platforms like Illumina. The basic principle is to mutagenise sample to actually remove

Lutz Froenicke on Twitter: "Longas Technologies is going to offer a new synthetic long read technology: MorphoSeq. The scheme looks somewhat similar to the Loop Genomics approach and sequences fragments up to
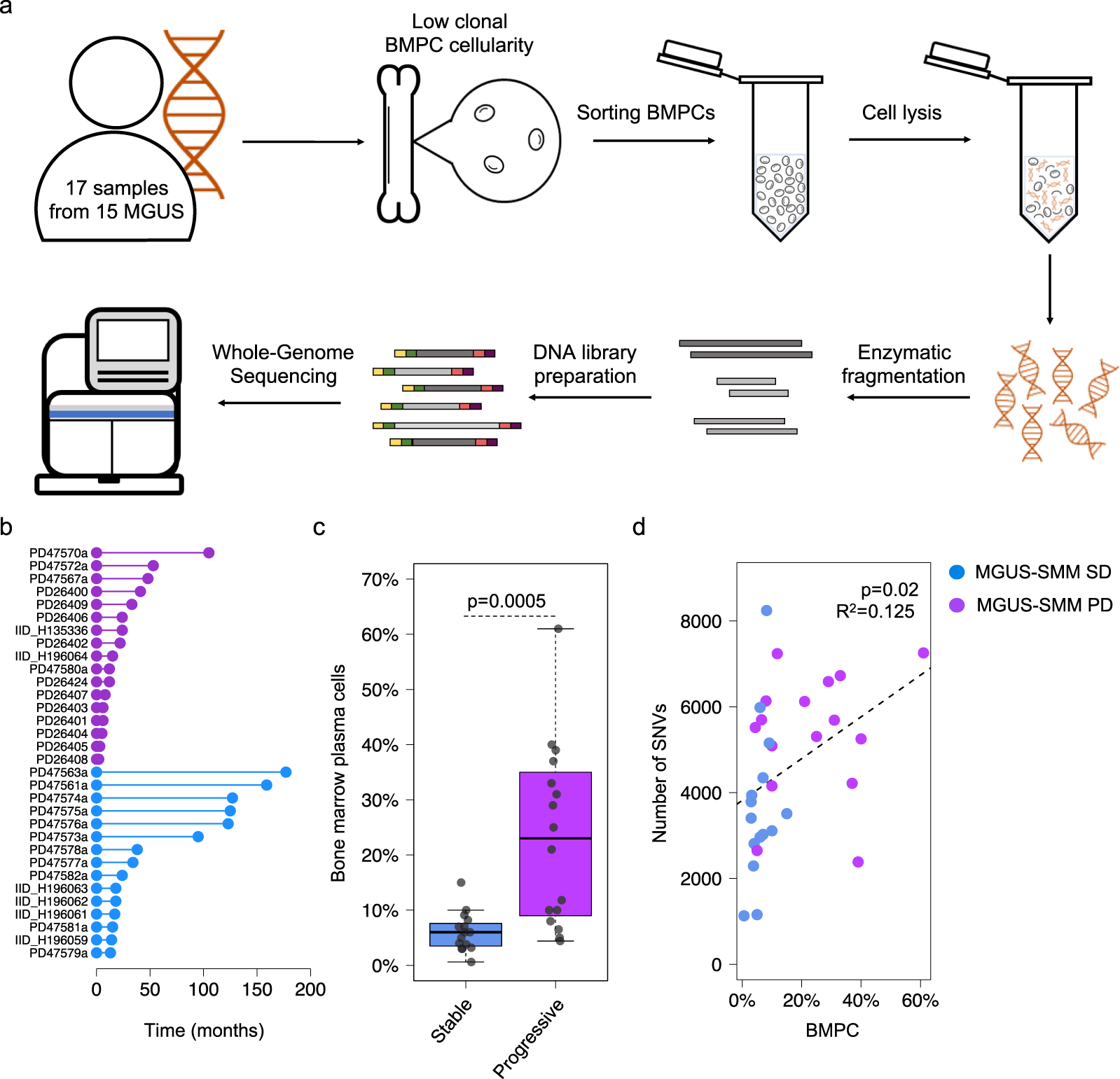
Whole-genome sequencing reveals progressive versus stable myeloma precursor conditions as two distinct entities | Nature Communications