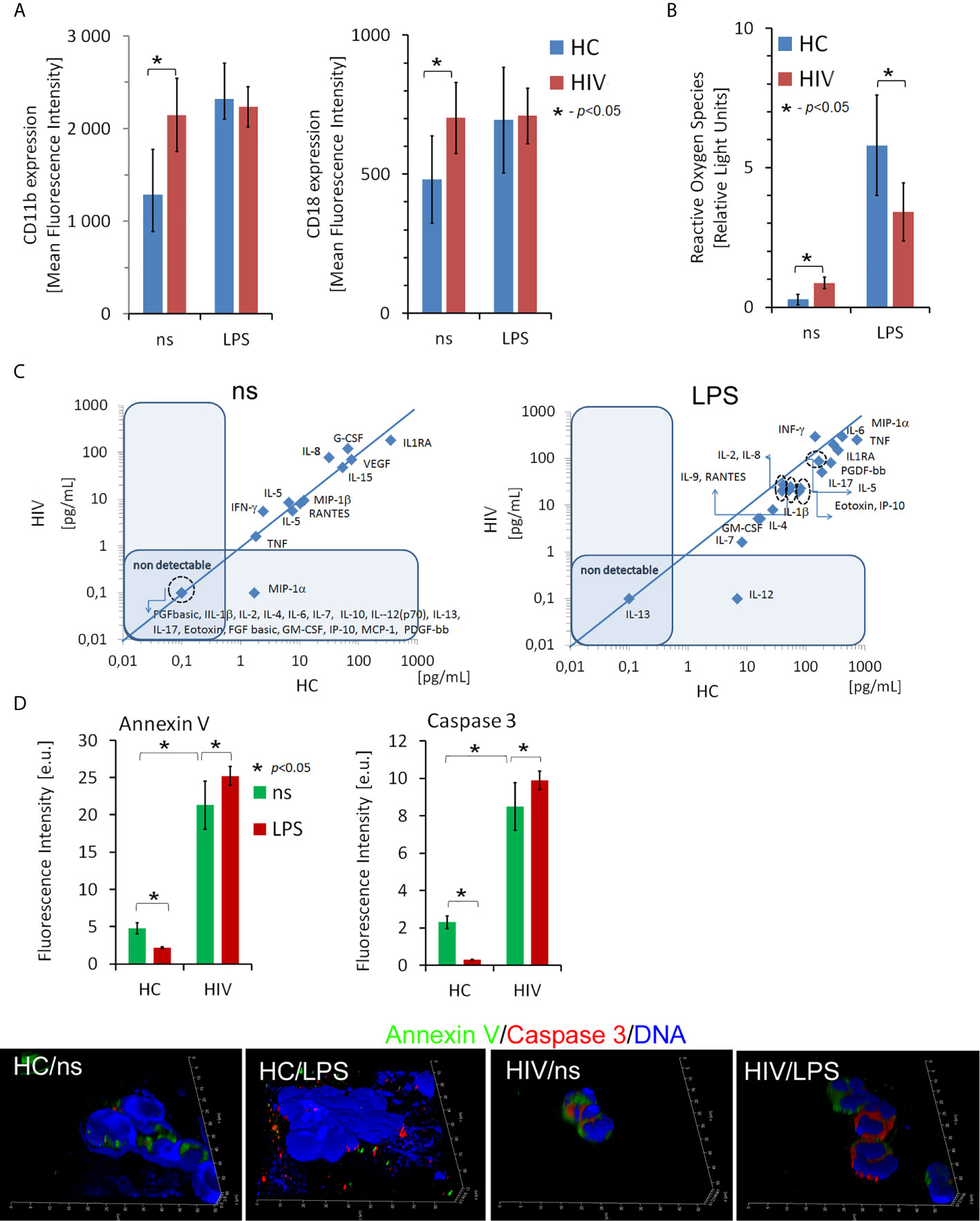
Frontiers | H3K4me3 Histone ChIP-Seq Analysis Reveals Molecular Mechanisms Responsible for Neutrophil Dysfunction in HIV-Infected Individuals

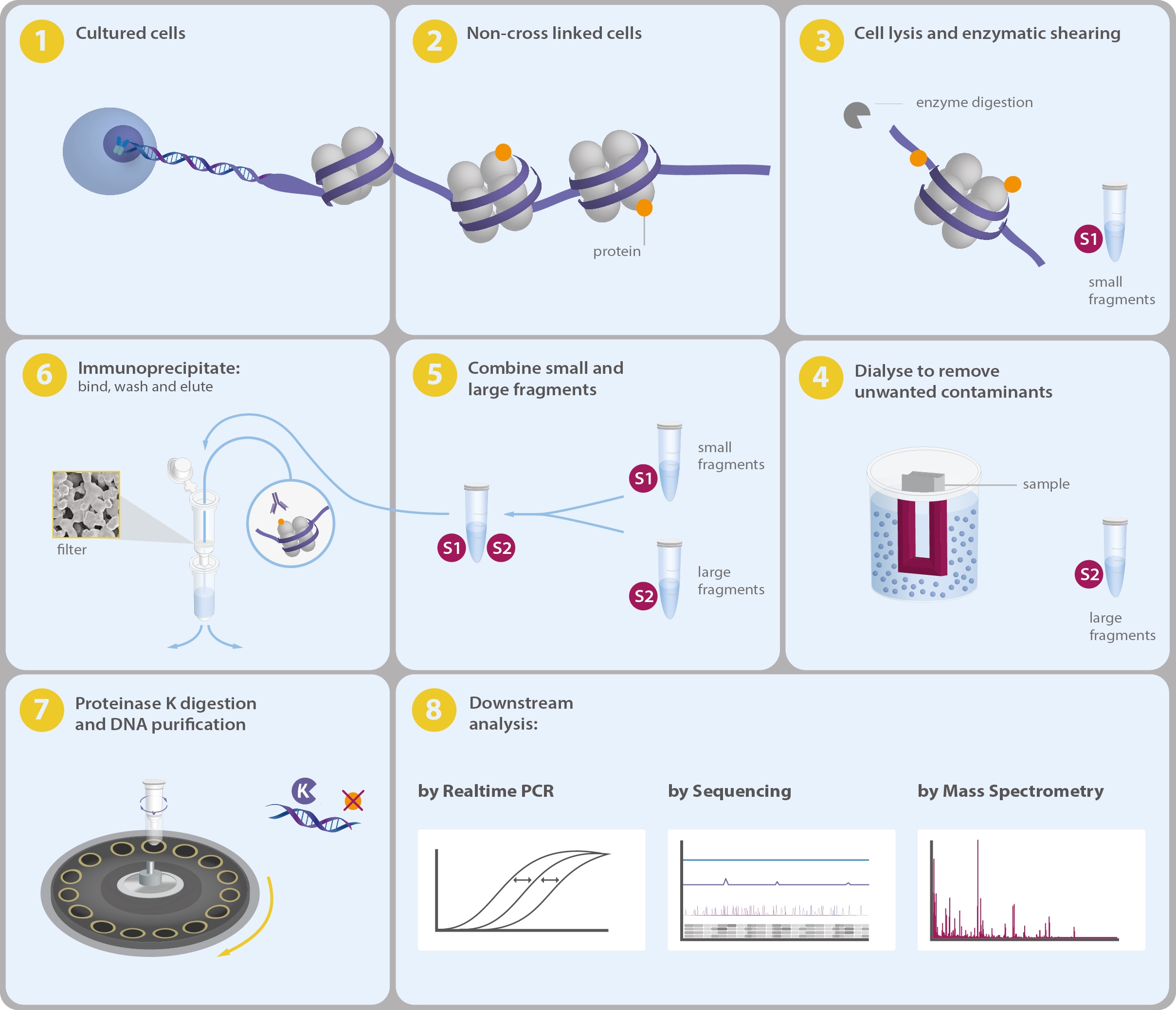
An optimized protocol for rapid, sensitive and robust on-bead ChIP-seq from primary cells - ScienceDirect

Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins | PNAS