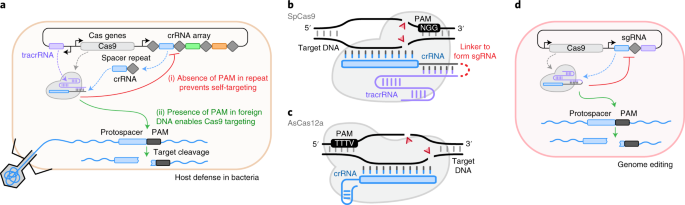
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols
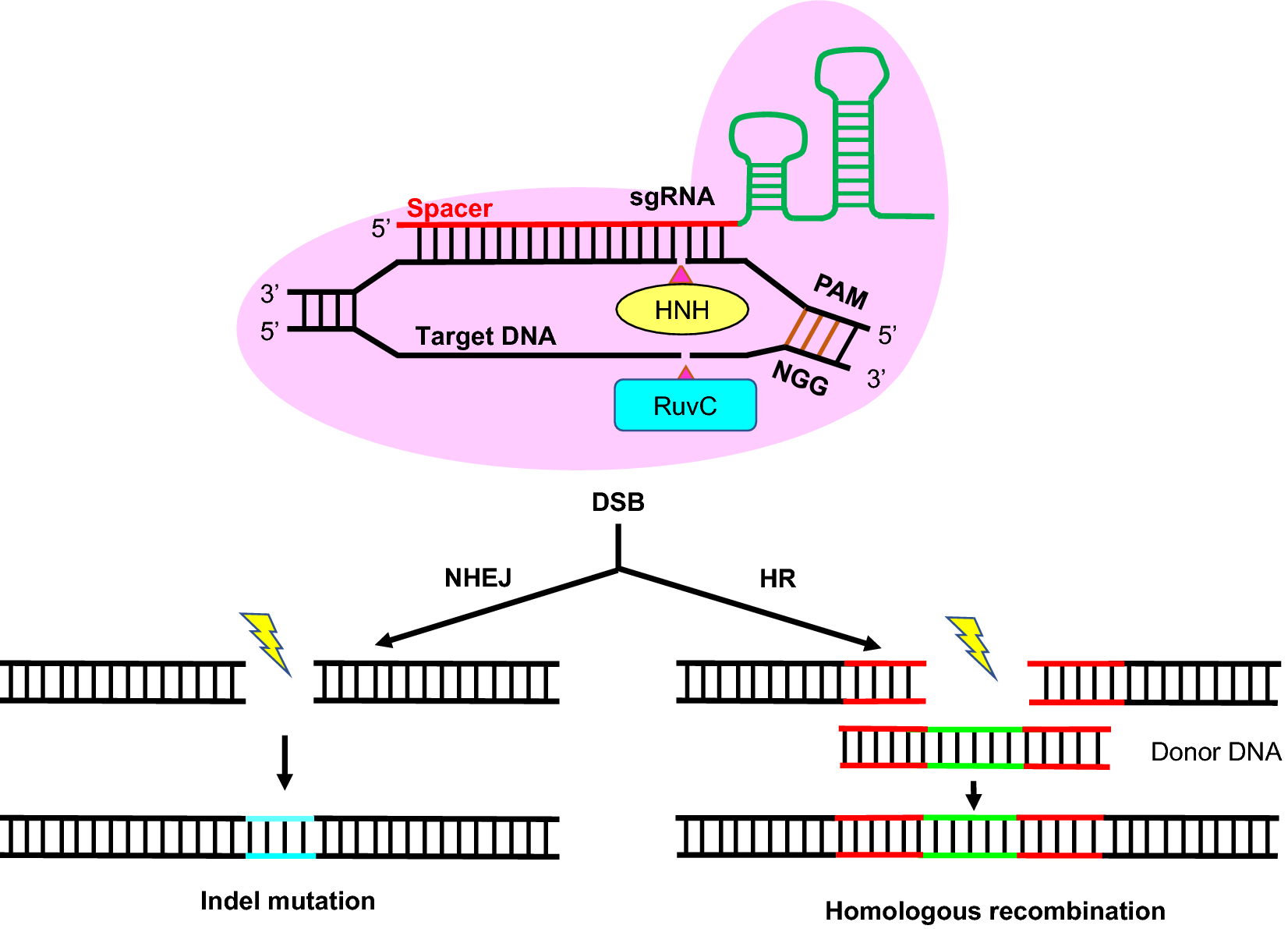
CRISPR-mediated genome editing in non-conventional yeasts for biotechnological applications | Microbial Cell Factories | Full Text
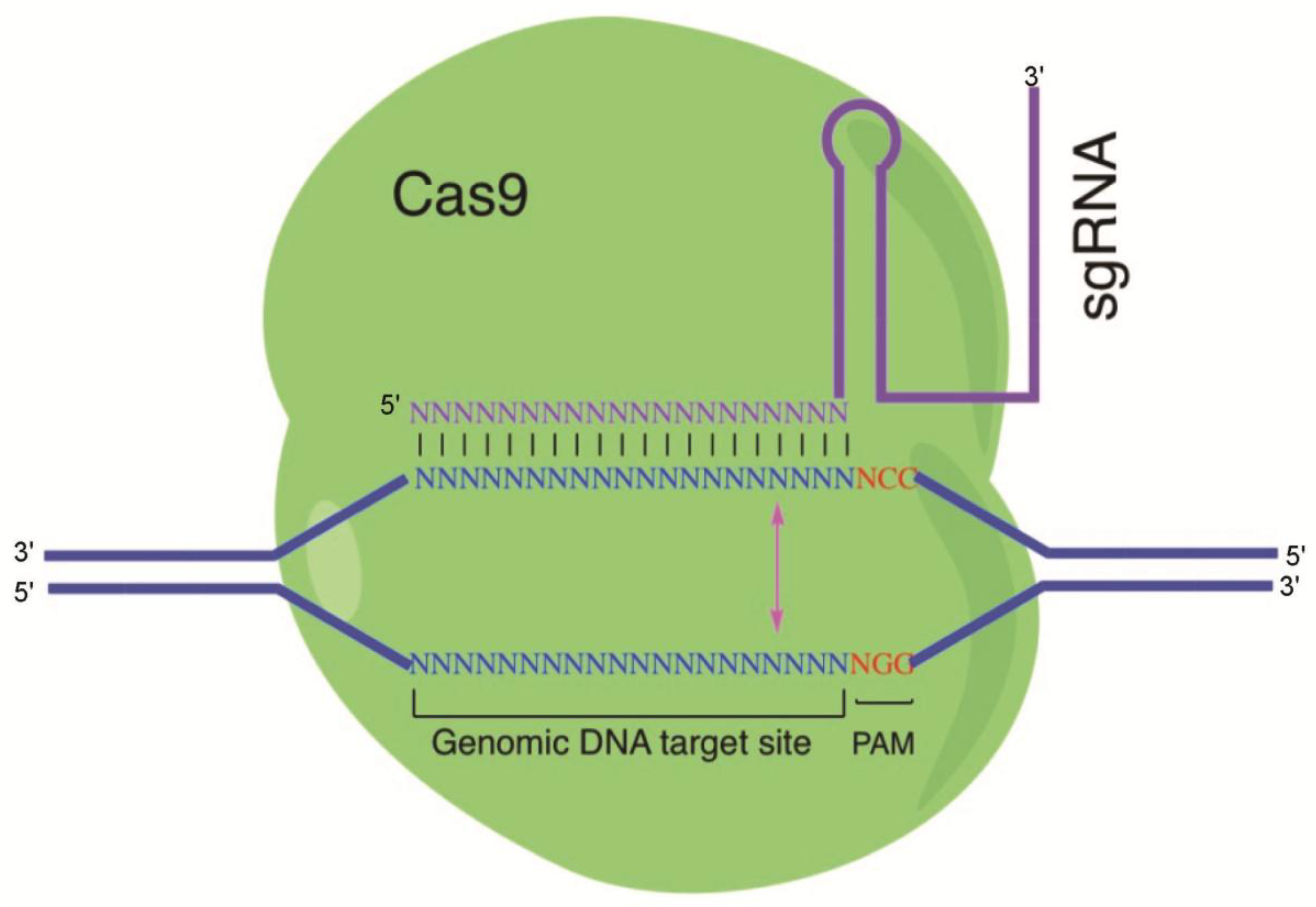
Viruses | Free Full-Text | Potential Application of the CRISPR/Cas9 System against Herpesvirus Infections

Comprehensive PAM prediction for CRISPR-Cas systems reveals evidence for spacer sharing, preferred strand targeting and conserved links with CRISPR repeats | bioRxiv
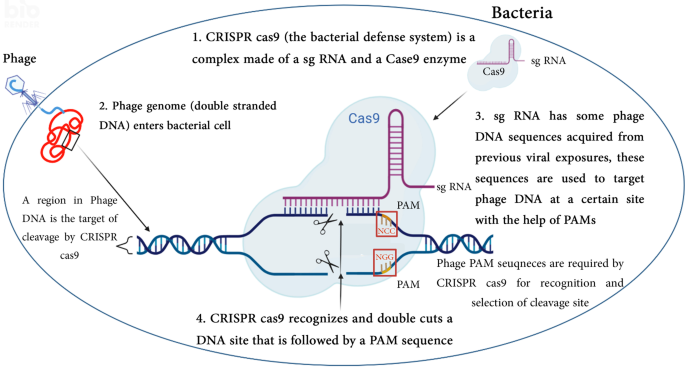
Bio-informatic analysis of CRISPR protospacer adjacent motifs (PAMs) in T4 genome | BMC Genomic Data | Full Text





.png?sfvrsn=58c63b07_0)